Research Articles
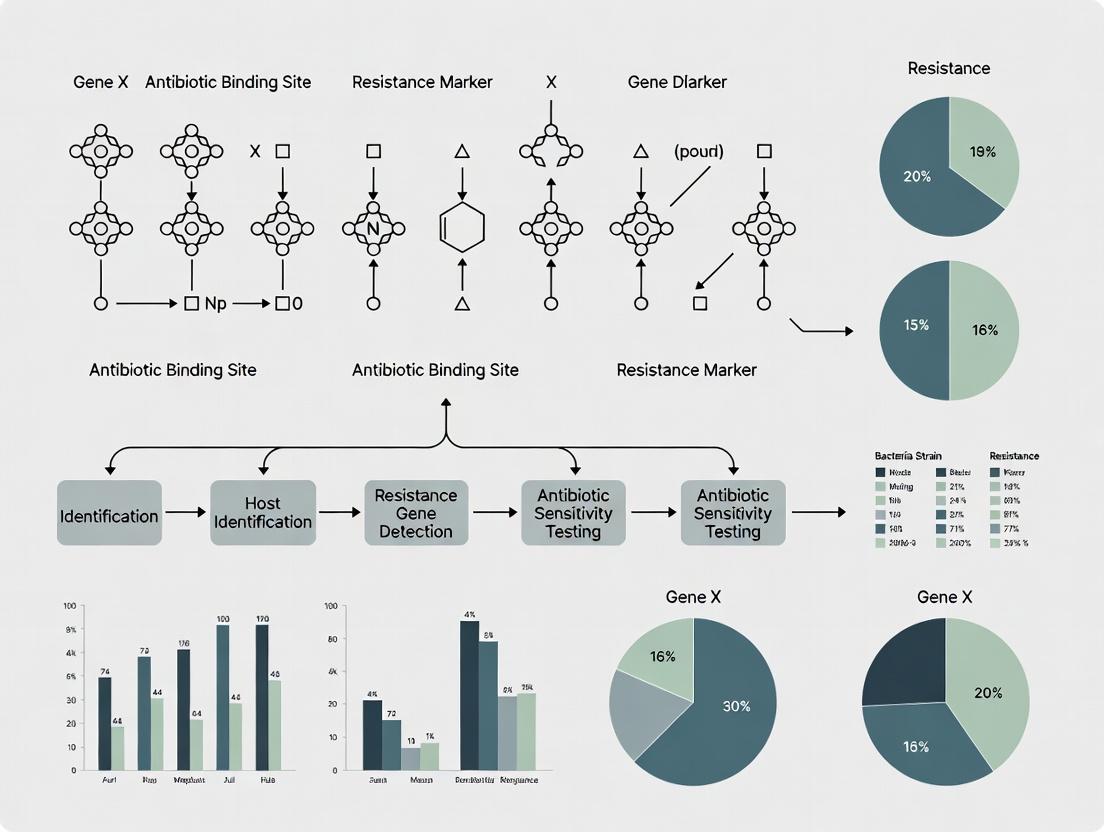
Binning Tools Benchmark 2024: Best Methods for Identifying Antibiotic Resistance Hosts in Metagenomic Data
This article provides a comprehensive guide and benchmark for bioinformatics tools used in binning metagenomic sequences to identify hosts of antibiotic resistance genes (ARGs).
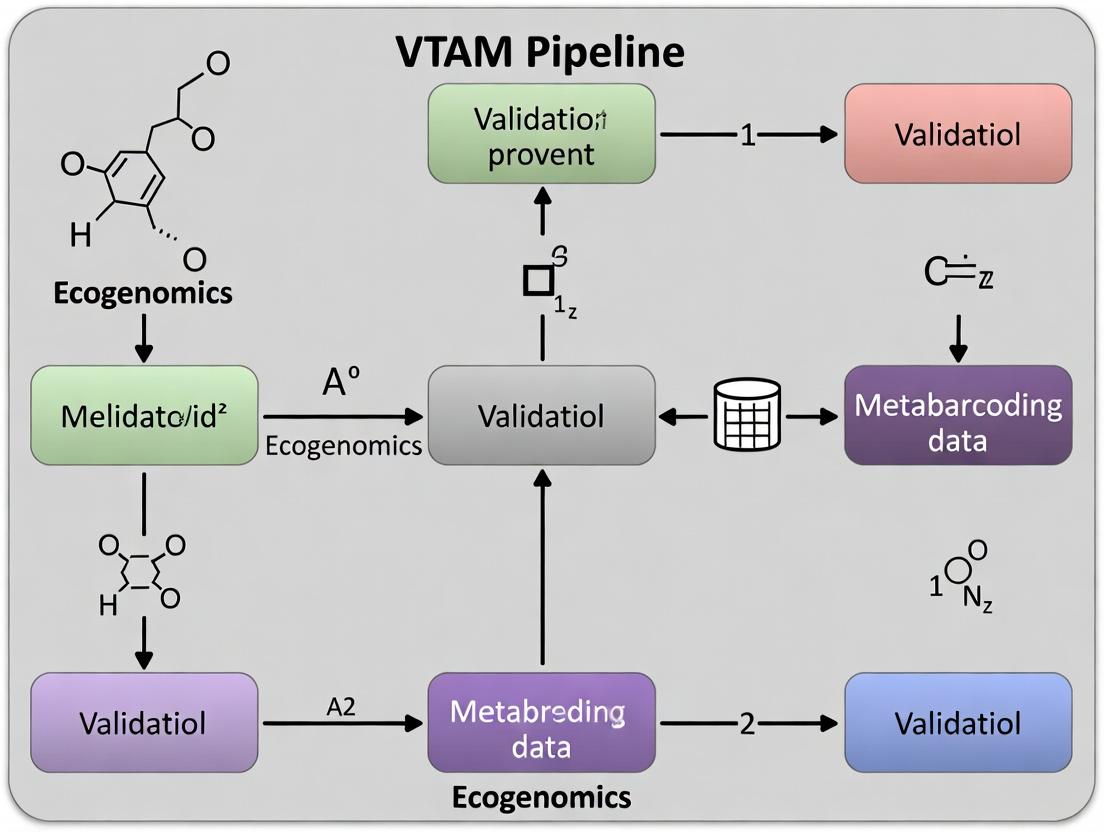
VTAM Pipeline: A Step-by-Step Guide to Validating Metabarcoding Data for Biomedical Research
This article provides a comprehensive guide to the VTAM (Validation, Taxonomic Assignment, and Analysis of Metabarcoding data) pipeline, a specialized tool for rigorous validation of amplicon sequence variants (ASVs) in...
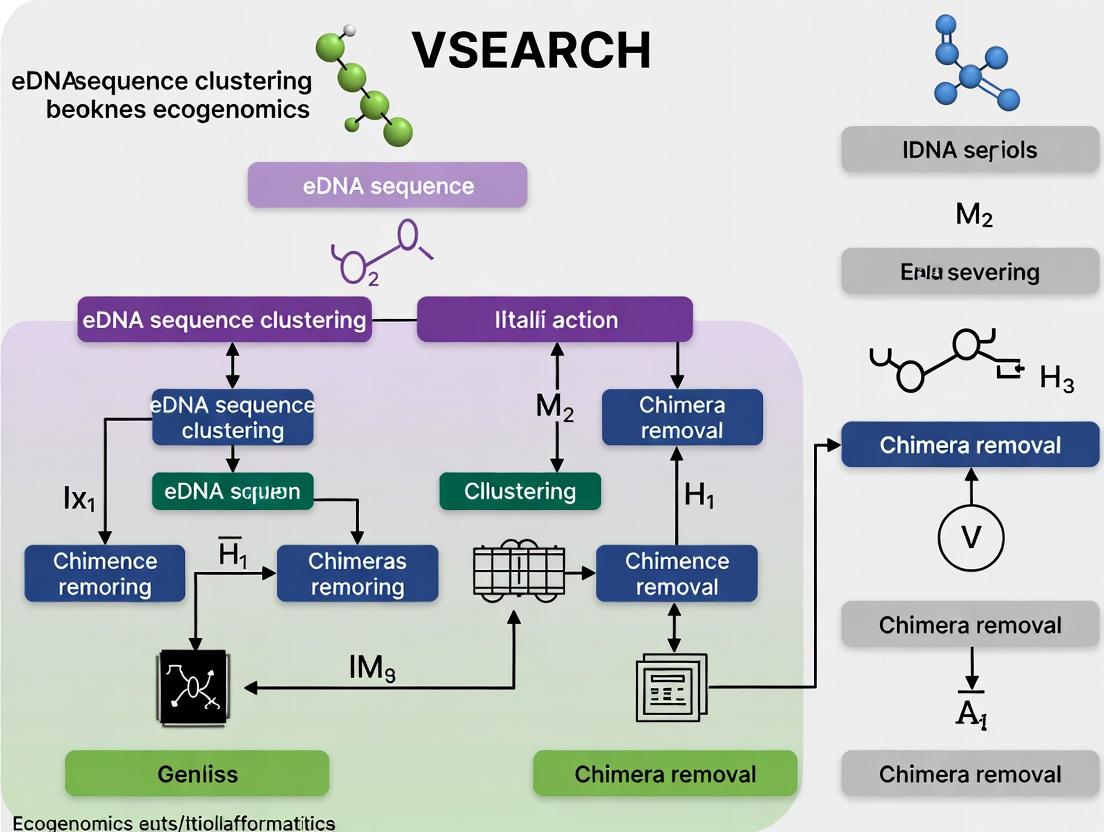
VSEARCH for eDNA Analysis: A Complete Guide to Sequence Clustering, Chimera Removal, and Bioinformatic Workflows
This comprehensive guide explores the critical role of VSEARCH in environmental DNA (eDNA) analysis for researchers and biopharma professionals.
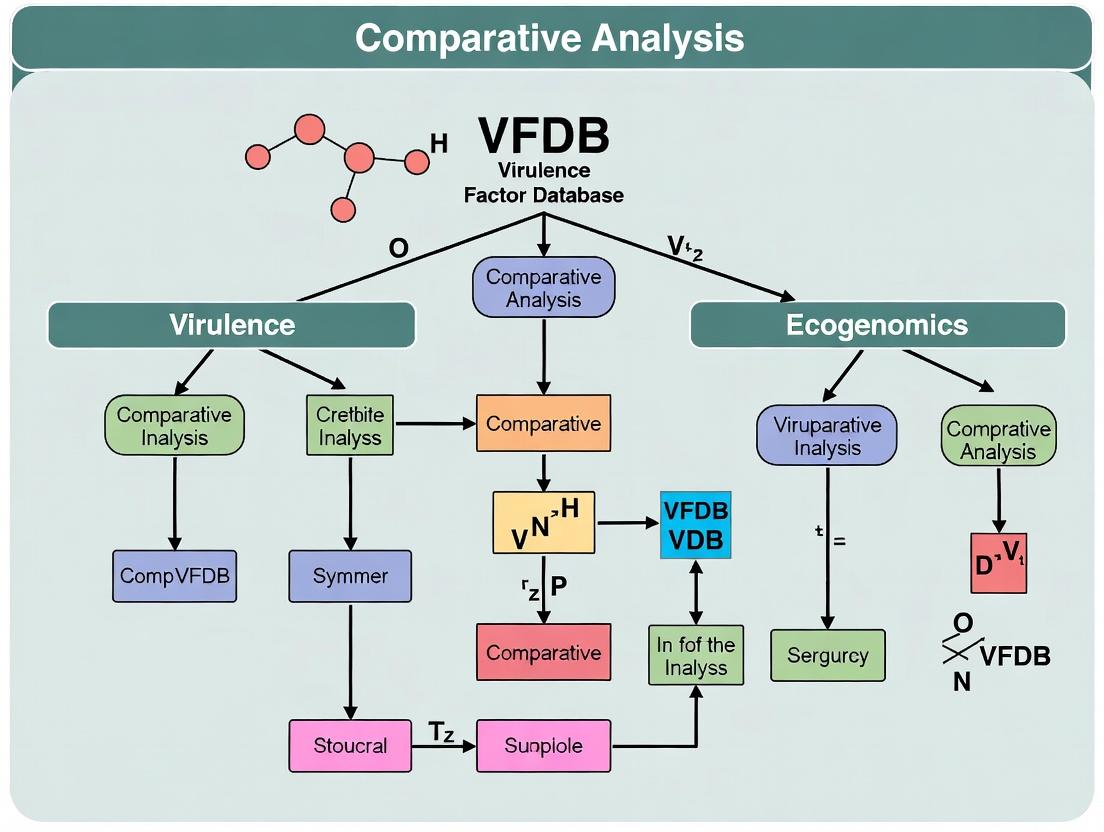
Mastering VFDB for Comparative Pathogenomics: A Researcher's Guide to Analyzing Virulence Factors
This article provides a comprehensive guide for researchers and drug development professionals on utilizing the Virulence Factor Database (VFDB) for robust comparative genomic analysis.
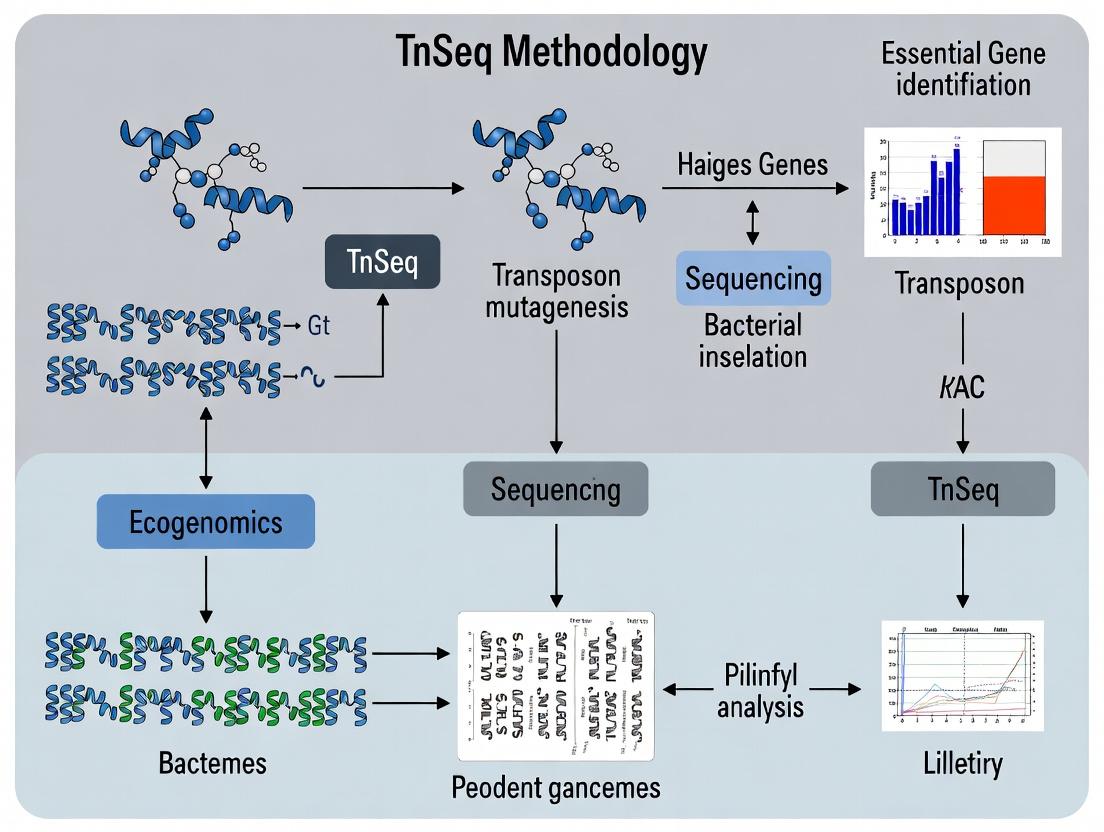
Essential Genes for Infection: A Complete Guide to TnSeq in Bacterial Pathogenesis Research
This article provides a comprehensive guide for researchers and drug development professionals on using Transposon Sequencing (TnSeq) to identify bacterial genes essential for infection.
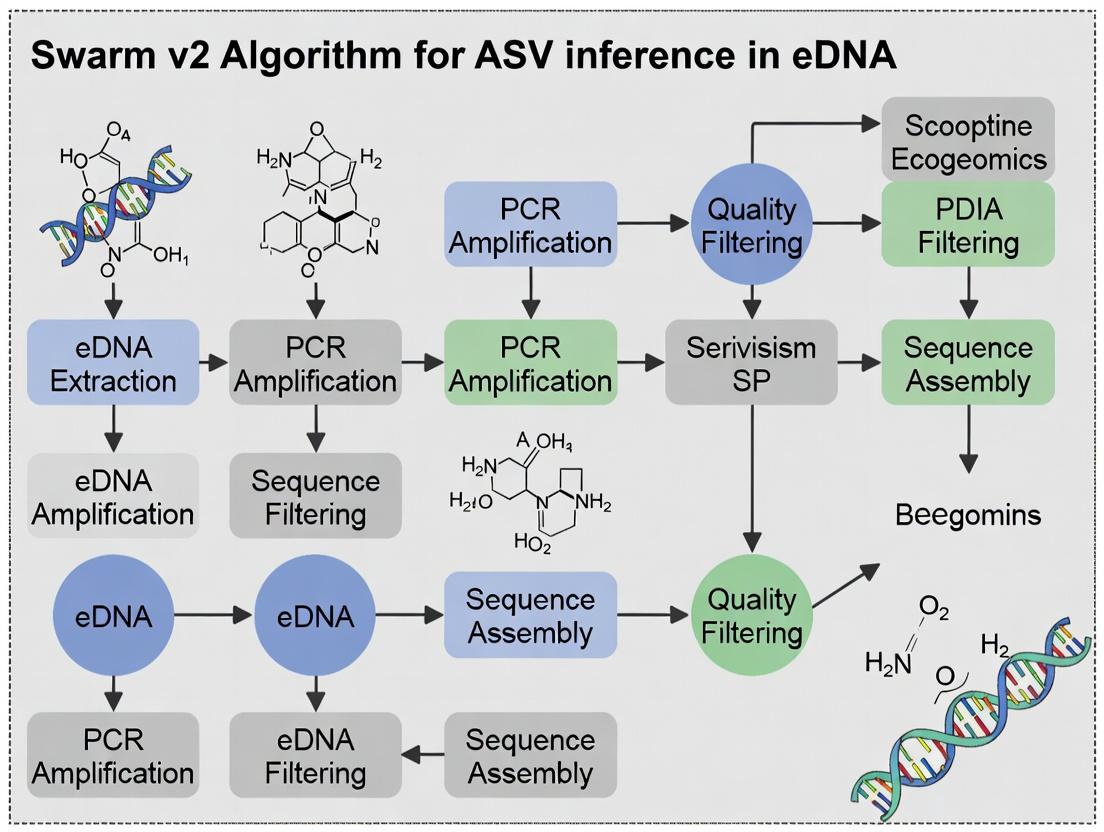
Unlocking Microbial Diversity: A Guide to Swarm v2 for Accurate ASV Inference in eDNA Research
This article provides a comprehensive overview of the Swarm v2 algorithm for Amplicon Sequence Variant (ASV) inference in environmental DNA (eDNA) studies, tailored for researchers and drug development professionals.
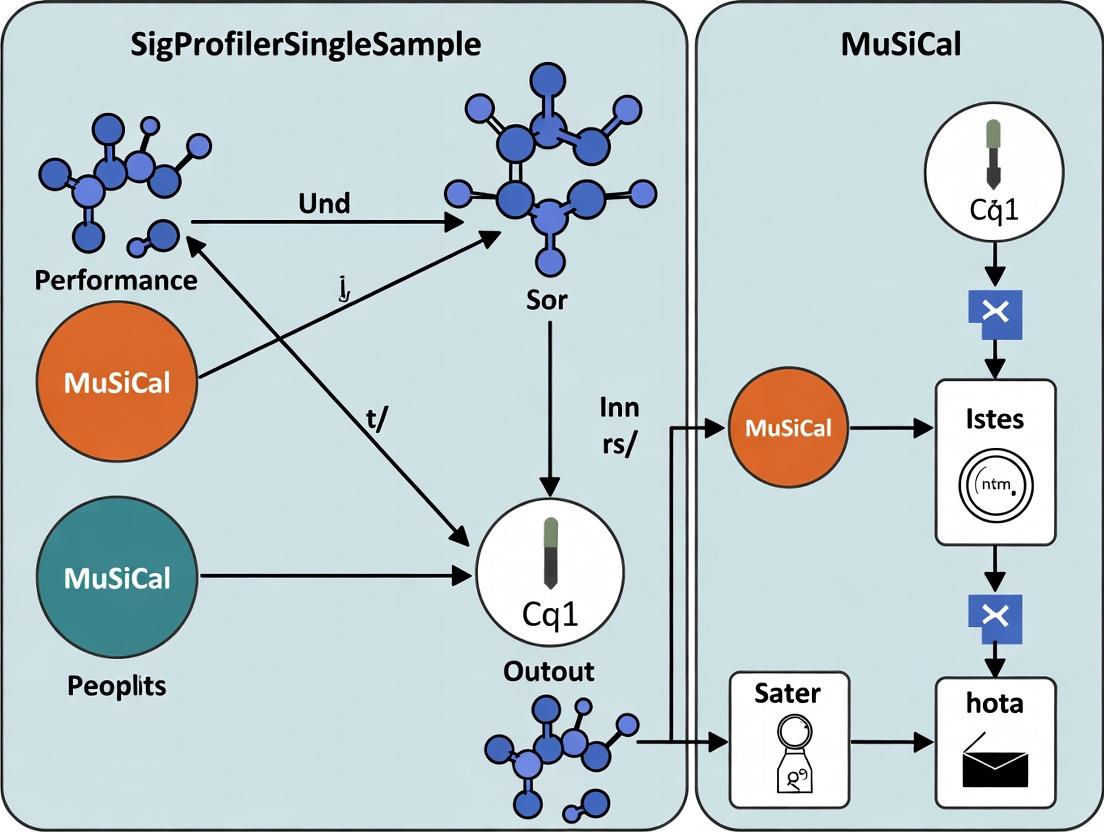
SigProfilerSingleSample vs MuSiCal: A Comprehensive 2024 Performance Evaluation for Cancer Mutational Signature Analysis
This article provides a detailed, current evaluation of two leading tools for mutational signature analysis: SigProfilerSingleSample (SPSS) and MuSiCal.
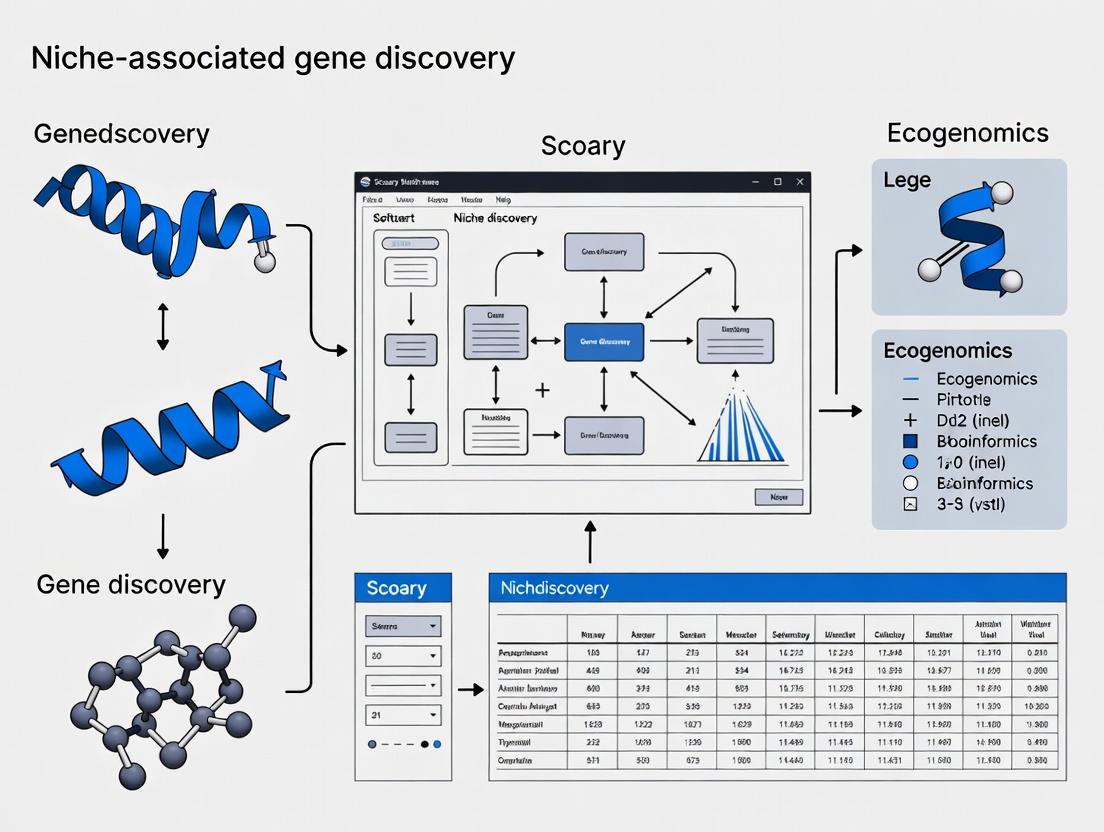
Scoary: The Essential Guide to Discovering Niche-Specific Genes in Microbial Genomes for Biomedical Research
This comprehensive guide explores Scoary, a powerful software tool for identifying genes associated with specific microbial traits or ecological niches using pan-genome-wide association studies (pan-GWAS).
Scoary: A Comprehensive Guide to Identifying Adaptive Genes in Microbial and Cancer Genomics
This guide provides a detailed exploration of Scoary, a powerful software tool for identifying genes associated with microbial phenotypes, such as antibiotic resistance, virulence, and host adaptation.
Scoary: A Comprehensive Guide to Identifying Niche-Associated Genes for Biomarker and Drug Target Discovery
This guide provides researchers, scientists, and drug development professionals with a complete framework for using Scoary, a powerful tool for identifying genes associated with microbial phenotypic niches from pan-genome data.