Hi-C Proximity Ligation for Phage-Host Identification: A Complete Guide for Research and Drug Development
This article provides a comprehensive guide to Hi-C proximity ligation for linking bacteriophages to their bacterial hosts, a critical step in phage therapy and microbiome research.
Hi-C Proximity Ligation for Phage-Host Identification: A Complete Guide for Research and Drug Development
Abstract
This article provides a comprehensive guide to Hi-C proximity ligation for linking bacteriophages to their bacterial hosts, a critical step in phage therapy and microbiome research. We explore the foundational principles of chromatin conformation capture adapted for virus-host interactions, detail step-by-step methodologies from sample preparation to data analysis, address common troubleshooting and optimization challenges, and validate the technique against alternative methods like metagenomics and microfluidics. Aimed at researchers and drug development professionals, this resource synthesizes current best practices to enable precise phage-host pairing for therapeutic discovery and ecological studies.
Decoding the Link: The Science Behind Hi-C for Phage-Host Interaction Mapping
Application Notes
Linking bacteriophages to their bacterial hosts is a critical challenge in viral ecology, microbiome research, and therapeutic development. The inability to culture most environmental bacteria (~99%) has historically obscured phage-host relationships. Hi-C proximity ligation methodology directly addresses this by capturing physical interactions between phage and host DNA within intact cells, enabling high-throughput, culture-independent linking. This approach is foundational for constructing accurate ecological networks and for rationally selecting phages for precision therapies against antibiotic-resistant pathogens.
Table 1: Comparison of Phage-Host Linking Methodologies
| Method | Principle | Throughput | Culture Requirement | Key Limitation | Typical Linking Accuracy |
|---|---|---|---|---|---|
| Hi-C Proximity Ligation | Captures chromatin contacts in situ | High (Metagenome-wide) | No | Requires high sequencing depth | >90% for dominant species |
| Viral Tagging (FACS) | Fluorescence-labeled phages bind hosts | Low | Yes (for hosts) | Limited to culturable hosts | ~95% for cultured pairs |
| CRISPR Spacer Analysis | Bioinformatic match of spacers to phages | Computational/High | No | Indirect evidence; historical links | Variable, high false negatives |
| Metagenomic Co-occurrence | Correlation of abundances across samples | Computational/High | No | Indirect; cannot distinguish infection | Low specificity |
Table 2: Hi-C Protocol Metrics and Outcomes (Representative Data)
| Parameter | Typical Value/Range | Impact on Results |
|---|---|---|
| Crosslinking Agent & Time | 3% Formaldehyde, 10-25 min | Under-fixing reduces contacts; over-fixing inhibits ligation. |
| Proximity Ligation Efficiency | 0.5-5% of total read pairs | Determines signal-to-noise ratio for linkage detection. |
| Sequencing Depth Requirement | 50-200M read pairs per metagenomic sample | Scales with community complexity and desired resolution. |
| Reported Linking Yield | 10-1000 phage-host links per sample (Marine/Soil) | Dependent on viral abundance and diversity. |
| Validation Rate (vs. culture) | 85-98% | Confirms high specificity of Hi-C links. |
Protocols
Detailed Protocol: Hi-C for Phage-Host Linking from Environmental Samples
Objective: To identify physical interactions between phage and bacterial host genomes within an uncultured microbial community.
Materials & Reagents
- Fixative: Formaldehyde (3% final concentration in buffer).
- Lysis Buffer: 50 mM Tris-HCl (pH 8.0), 50 mM NaCl, 1% SDS, plus protease inhibitors.
- Restriction Enzyme: 4-cutter (e.g., MboI, DpnII) or 6-cutter with frequent recognition in bacterial/viral genomes.
- Ligation Master Mix: T4 DNA Ligase, buffer, ATP, BSA, Triton X-100 (to quench SDS).
- DNA Cleanup: Solid-phase reversible immobilization (SPRI) beads.
- Quantification: Fluorometric dsDNA assay (e.g., Qubit).
- Sequencing: Illumina-compatible library prep kit, paired-end sequencing.
Procedure
Sample Fixation & Crosslinking:
- Collect biomass (e.g., from water, soil, or fecal sample) and resuspend in appropriate buffer.
- Add formaldehyde to 3% final concentration. Incubate at room temperature for 25 minutes with gentle rotation.
- Quench crosslinking by adding 2.5M glycine to a final concentration of 0.625M. Incubate 5 minutes at room temperature.
- Pellet cells, wash 2x with cold PBS.
Cell Lysis & Chromatin Digestion:
- Resuspend pellet in lysis buffer. Incubate at 65°C for 15 minutes.
- Dilute SDS concentration to <0.1% using 1% Triton X-100.
- Digest chromatin with 400U of restriction enzyme (e.g., MboI) overnight at 37°C with rotation.
Proximity Ligation & Crosslink Reversal:
- Dilinate digested DNA ends with biotinylated nucleotides using Klenow fragment.
- Under dilute conditions (to favor intra-molecular ligation), add T4 DNA Ligase and ligate for 4 hours at room temperature.
- Reverse crosslinks by adding Proteinase K and incubating at 65°C overnight.
- Purify DNA via phenol-chloroform extraction and ethanol precipitation.
Biotin Pull-down & Library Preparation:
- Shear DNA to ~500 bp fragments using a sonicator.
- Bind biotin-labeled ligation junctions to streptavidin-coated magnetic beads.
- On-bead, perform end-repair, A-tailing, and adapter ligation for Illumina sequencing.
- Perform a final PCR amplification (12-16 cycles) with index primers. Clean up with SPRI beads.
Sequencing & Bioinformatic Analysis:
- Sequence on an Illumina platform (minimum 2x150 bp, target 100-200M read pairs).
- Process data through a dedicated pipeline (e.g., hicstuff, Juicer, or metaHiC):
- Trim and map read pairs to a composite reference database of bacterial and viral genomes.
- Identify valid interaction pairs (ligation junctions).
- Statistically assign phage contigs to bacterial host genomes based on significant enrichment of contact frequency versus background.
Diagrams
Hi-C Phage-Host Linking Workflow
Molecular Basis of Hi-C Linking
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for Hi-C Phage-Host Linking
| Item | Function in Protocol | Key Considerations |
|---|---|---|
| Formaldehyde (37%) | Crosslinks phage and host DNA in situ within infected cells. | Fresh aliquots preferred; concentration and time must be optimized for sample type. |
| Frequent-Cutting Restriction Enzyme (e.g., MboI) | Digests crosslinked DNA to create ends for ligation. | Choose enzyme(s) with high frequency in expected bacterial/viral genomes (4-6 bp cutter). |
| Biotin-14-dATP/dCTP | Labels digested DNA ends for selective pull-down of ligation junctions. | Critical for enriching chimeric fragments over non-ligated background. |
| Streptavidin Magnetic Beads | Isolates biotinylated proximity ligation products post-sonication. | High binding capacity and low non-specific DNA retention are essential. |
| Phase Lock Gel Tubes | Facilitates clean phenol-chloroform extraction after crosslink reversal. | Maximizes recovery of high-molecular-weight, crosslinked DNA. |
| Comprehensive Reference Database (e.g., RefSeq, IMG/VR) | For mapping sequenced read pairs to bacterial and viral genomes. | Quality and completeness directly limit discovery; should include metagenome-assembled genomes (MAGs). |
| Bioinformatics Pipeline (e.g., metaHiC) | Processes sequencing data to identify statistically significant phage-host contacts. | Must handle metagenomic mapping, noise filtering, and statistical modeling (e.g., binomial test). |
Within the context of a broader thesis on using Hi-C proximity ligation for phage-host linking research, understanding the core biochemical principle is fundamental. Proximity ligation is a molecular biology technique that converts transient physical interactions between DNA segments into stable, sequenceable chimeric DNA molecules. This allows for the genome-wide mapping of chromosomal contacts and, in metagenomic applications, the identification of which phage DNA is physically associated with which host bacterial genome.
The Core Biochemical Principle
The principle rests on crosslinking, digestion, ligation, and purification. First, living cells are treated with formaldehyde, which creates covalent crosslinks between DNA and proteins, and, crucially, between DNA strands that are in close spatial proximity (typically < 10 nm). This "freezes" the 3D genomic architecture. The crosslinked DNA is then digested with a restriction enzyme, creating fragments with compatible sticky ends. Under dilute conditions that favor intramolecular ligation, these sticky ends are ligated. Critically, only DNA ends that are held in close proximity by crosslinks will be ligated together, creating "chimeric junctions." After reversing crosslinks and purifying the DNA, these chimeric fragments can be sequenced. The pairs of sequences that form the junction are inferred to have been in physical contact in the native cell.
Application in Phage-Host Linking
In phage-host research, this principle is applied to environmental or laboratory samples containing a mixture of bacteria and their viral predators (phages). Crosslinking captures both intra-genomic contacts and inter-genomic contacts, such as those between a prophage integrated into a bacterial chromosome or between an infecting phage genome and its host genome. Sequencing and bioinformatic analysis of the chimeric reads allow the assignment of phages to their specific microbial hosts based on the statistical enrichment of contact frequencies.
Table 1: Key Parameters in a Standard Hi-C/Proximity Ligation Protocol for Microbial Communities
| Parameter | Typical Value or Specification | Purpose/Rationale |
|---|---|---|
| Crosslinking Agent | 1-3% Formaldehyde | Fixes spatial proximity of DNA segments. |
| Crosslinking Time | 10-30 minutes (at room temp) | Balances efficient crosslinking with over-crosslinking. |
| Restriction Enzyme | 4-cutter (e.g., DpnII, MboI, HindIII) | Creates frequent fragments for high-resolution contact maps. |
| Ligation Condition | Dilute, Blunt-end after fill-in | Favors ligation of crosslinked, proximate ends over random ligation. |
| Sequencing Depth | 50-200 million read pairs (microbial) | Sufficient to detect lower-frequency inter-genomic contacts. |
| Valid Chimeric Read Rate | 10-30% of total reads | Metric for protocol efficiency; depends on sample and prep. |
| Crosslink Reversal | Incubation at 65°C with Proteinase K | Cleaves formaldehyde crosslinks to purify DNA. |
Table 2: Bioinformatic Output Metrics from Phage-Host Hi-C Analysis
| Metric | Description | Implication for Phage-Host Linking |
|---|---|---|
| Contact Frequency | Raw count of chimeric reads linking two genomic loci. | Direct measure of interaction strength. |
| Statistical Significance (p-value) | Probability contact frequency occurs by chance. | Identifies confident, non-random phage-host associations. |
| Interaction Distance | Genomic distance from contact point to host integration site (for prophages). | Distinguishes integrated prophages from transient infections. |
| Host Range Breadth | Number of distinct host species linked to a single phage. | Informs on phage specificity (narrow vs. broad host range). |
Experimental Protocol: Hi-C for Phage-Host Linking from Environmental Samples
Protocol: Metagenomic Hi-C for Phage-Host Identification
I. Sample Collection and Crosslinking
- Collect biomass from environmental sample (e.g., water, soil, gut content) and resuspend in PBS.
- Add formaldehyde to a final concentration of 2% and incubate at room temperature for 25 minutes with gentle rotation.
- Quench the crosslinking reaction by adding glycine to a final concentration of 0.2 M and incubate for 5 minutes.
- Pellet cells by centrifugation, wash twice with cold PBS, and flash-freeze pellet for storage at -80°C or proceed.
II. Cell Lysis and Chromatin Digestion
- Resuspend crosslinked pellet in 1x appropriate restriction enzyme buffer with 0.1% SDS. Incubate at 37°C for 1 hour with shaking to lyse cells and expose chromatin.
- Quench SDS by adding Triton X-100 to 1%.
- Add 200-400 units of a frequent-cutter restriction enzyme (e.g., MboI or DpnII). Incubate at 37°C overnight with rotation.
III. Fill-in and Proximity Ligation
- Fill in restriction overhangs and incorporate biotinylated nucleotides. Use Klenow fragment (exo-) in the presence of dATP, dGTP, dTTP, and biotin-14-dCTP. Incubate at 37°C for 1.5 hours.
- Dilute the reaction mix with ligation buffer to ~4 ml to favor intramolecular ligation.
- Add T4 DNA Ligase and incubate at 16°C for 6 hours.
IV. Crosslink Reversal and DNA Purification
- Reverse crosslinks by adding Proteinase K and incubating at 65°C overnight.
- Purify DNA by phenol:chloroform extraction and ethanol precipitation.
- Shear purified DNA to ~300-500 bp fragments using a sonicator.
- Perform size selection to remove very small fragments.
V. Biotin Pulldown and Library Preparation
- Bind biotin-labeled chimeric junctions to streptavidin-coated magnetic beads.
- Wash beads thoroughly to remove non-biotinylated DNA.
- On-bead, perform end-repair, A-tailing, and adapter ligation for Illumina sequencing.
- Perform a final PCR amplification (with limited cycles) to add full indexing.
- Purify the final library and quantify via qPCR for sequencing on an Illumina platform.
Visualizations
Diagram: Hi-C Workflow for Phage-Host Linking
Diagram: Molecular Steps of Proximity Ligation
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for Hi-C-based Phage-Host Linking
| Item | Function in Protocol | Key Considerations |
|---|---|---|
| Formaldehyde (37%) | In-situ crosslinking agent to fix DNA-protein and DNA-DNA contacts. | Use fresh, molecular biology grade. Quenching with glycine is critical. |
| Frequent-Cutter Restriction Enzyme (e.g., DpnII) | Digests crosslinked DNA to create ligatable ends; determines resolution. | Choose enzyme compatible with expected G/C content of community DNA. |
| Biotin-14-dCTP | Biotinylated nucleotide used in fill-in reaction to label ligation junctions. | Allows for stringent streptavidin-based enrichment of chimeric fragments. |
| T4 DNA Ligase | Catalyzes the ligation of crosslink-proximal DNA ends. | High-concentration enzyme used under dilute conditions. |
| Streptavidin Magnetic Beads | Solid-phase support for affinity purification of biotinylated chimeric DNA. | High binding capacity and low non-specific DNA binding are essential. |
| Proteinase K | Protease that aids in reversing formaldehyde crosslinks during DNA purification. | Requires long incubation at high temperature (65°C). |
| Klenow Fragment (exo-) | DNA polymerase for fill-in of sticky ends; lacks exonuclease activity. | Ensures efficient incorporation of biotin-dCTP. |
| Size Selection Beads (SPRI) | For clean-up and size selection of DNA after shearing and library prep. | Critical for removing small fragments and adapter dimers. |
| Paired-End Sequencing Kit (Illumina) | Generates sequence data from both ends of the chimeric fragment. | Allows mapping of each read pair to potentially distinct genomes. |
| Bioinformatics Pipeline (e.g., HiC-Pro, distiller) | Processes raw sequences, maps reads, filters artifacts, generates contact matrices. | Must be adapted for metagenomic mode to handle multiple genomes. |
This application note details the adaptation of chromosome conformation capture (Hi-C) technology from its origins in mammalian 3D genomics to its groundbreaking application in linking bacteriophages to their bacterial hosts. Within the broader thesis on Hi-C proximity ligation for phage host linking, this document provides the essential protocols and data analysis frameworks required to successfully apply this tool in microbiome and therapeutic discovery research.
Table 1: Comparison of Hi-C Protocol Parameters Across Biological Systems
| Parameter | Mammalian Chromosomes (Original) | Microbial Communities (Adapted) | Phage-Host Linking (Specialized) |
|---|---|---|---|
| Crosslinking Agent | 1-3% Formaldehyde | 3% Formaldehyde + 1% DSG (disuccinimidyl glutarate) | 3% Formaldehyde |
| Crosslinking Time | 10-30 min | 30-45 min | 20-30 min |
| Cell Lysis Method | Detergent-based (NP-40, SDS) | Enzymatic (lysozyme) + Detergent | Enzymatic (lysozyme, mutanolysin) + Detergent |
| Ligation Strategy | Biotin-labeled blunt-end ligation | Biotin-labeled blunt-end ligation | Biotin-labeled blunt-end ligation |
| Typical Sequencing Depth | 1-5 Billion reads | 50-200 Million reads | 20-100 Million reads |
| Key Analytical Output | TADs, Compartments, Loops | Species deconvolution, plasmids | Phage-host contact frequency |
Table 2: Representative Hi-C Phage-Host Linking Results (Meta-Analysis)
| Study Sample Type | % of Phages Linked to Host | Common Linked Host Genera | Detection Limit (Community Complexity) |
|---|---|---|---|
| Human Gut Microbiome | 40-60% | Bacteroides, Faecalibacterium, Escherichia | Up to 100+ species |
| Marine Microbial Community | 20-35% | Synechococcus, Pelagibacter | Up to 50+ species |
| Soil Microbiome | 15-30% | Pseudomonas, Bacillus, Streptomyces | Up to 150+ species |
| Enriched Lab Culture | >95% | Target-specific | <10 species |
Experimental Protocols
Protocol A: Hi-C for Phage-Host Linking from Complex Communities
I. Sample Fixation and Crosslinking
- Collect microbial biomass (e.g., 0.5g stool, 50ml filtered seawater) into 5 ml of cold PBS.
- Add formaldehyde to a final concentration of 3% (v/v). Mix thoroughly.
- Incubate at room temperature for 30 minutes with gentle rotation.
- Quench crosslinking by adding 2.5M glycine to a final concentration of 0.2M. Incubate for 10 min at RT.
- Pellet cells at 8,000 x g for 5 min at 4°C. Wash pellet 2x with cold PBS.
II. Cell Lysis and Chromatin Digestion
- Resuspend pellet in 1ml ice-cold lysis buffer (50 mM Tris-HCl pH 8.0, 50 mM NaCl, 1% SDS, 1x protease inhibitor).
- Add 10 µl of 100 mg/ml lysozyme. Incubate 30 min at 37°C with gentle mixing.
- Quench SDS by adding 10% Triton X-100 to a final concentration of 1%.
- Add 100 U of DpnII or HindIII restriction enzyme. Incubate overnight at 37°C with rotation. Heat-inactivate enzyme as per manufacturer's instructions.
III. Proximity Ligation and DNA Purification
- Fill the digested lysate to 7 ml with ligation buffer (final concentration: 66 mM Tris-HCl pH 7.5, 5 mM NaCl, 5 mM MgCl2, 1% Triton X-100).
- Add 50 µl of 10 mg/ml BSA and 1,000 U of T4 DNA Ligase.
- Incubate for 4 hours at 16°C, followed by 30 min at room temperature.
- Reverse crosslinks by adding Proteinase K to 0.4 mg/ml and incubating overnight at 65°C.
- Purify DNA using standard phenol-chloroform extraction and ethanol precipitation. Resuspend in 100 µl TE buffer.
IV. Biotin Removal and Library Preparation
- Treat purified DNA with 5 U of T4 DNA Polymerase (in the absence of nucleotides) for 4 hours at 20°C to remove biotin from unligated ends.
- Shear DNA to ~300-500 bp using a focused ultrasonicator.
- Perform size selection using SPRI beads. Isolate biotinylated ligation junctions using streptavidin-coated magnetic beads.
- Perform on-bead library preparation for Illumina sequencing: end-repair, A-tailing, adapter ligation, and PCR amplification (12-15 cycles).
- Sequence on an Illumina platform using 2x150 bp paired-end chemistry.
Protocol B: In Silico Analysis Pipeline for Phage-Host Detection
- Preprocessing: Trim adapters and low-quality bases with Trimmomatic. Map paired-end reads to a combined reference database (bacterial genomes + viral genomes/contigs) using Bowtie2 with
--very-sensitiveand--no-discordantflags. - Pair Parsing: Parse aligned read pairs using a custom script or tools like HiC-Pro. Valid pairs are defined as two reads mapping to different restriction fragments.
- Contact Matrix Generation: Generate contact matrices for each sample, organizing by genome.
- Host Linking: Identify phage-host links by extracting trans contacts where one read maps to a viral contig and its mate maps to a bacterial genome. Apply a statistical filter (e.g., must be ≥5 unique read-pairs, and significant over background by binomial test).
- Visualization: Generate Circos plots or network graphs to visualize specific phage-host interaction networks.
Visualization Diagrams
Title: Hi-C Phage-Host Linking Workflow
Title: Molecular to In Silico Phage Host Detection
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for Hi-C Phage-Host Linking
| Reagent/Material | Function & Rationale | Example Product/Catalog |
|---|---|---|
| Disuccinimidyl Glutarate (DSG) | Membrane-permeable protein-protein crosslinker; enhances fixation of phage particles to host cell surfaces in complex samples. | Thermo Fisher, #20593 |
| Formaldehyde (37%) | Primary crosslinker for DNA-protein and protein-protein interactions; preserves in vivo chromatin and phage attachment structures. | MilliporeSigma, #252549 |
| HindIII or DpnII | Frequent-cutter restriction enzymes; used to digest crosslinked DNA to increase resolution of ligation junctions. | NEB, #R0104S (HindIII) |
| Biotin-14-dATP | Labels fragment ends during fill-in reaction; enables selective capture of ligated junctions via streptavidin beads. | Jena Bioscience, #NU-835-BIO14 |
| T4 DNA Ligase | Catalyzes intra- and inter-molecular ligation of crosslinked, digested DNA fragments; forms the chimeric junctions for sequencing. | NEB, #M0202S |
| Streptavidin C1 Beads | Magnetic beads for high-efficiency capture of biotinylated ligation junctions; critical for enriching for informative reads. | Invitrogen, #65001 |
| Protease K | Digests proteins to reverse formaldehyde crosslinks after ligation; releases DNA for purification and downstream processing. | Thermo Fisher, #AM2546 |
| Phage & Host Genomic DBs | Curated reference databases (e.g., NCBI Virus, GTDB) essential for accurate read mapping and host assignment. | NCBI, IMG/VR, GTDB |
Application Notes
In the context of phage therapy and microbiome research, accurately linking bacteriophages to their bacterial hosts is fundamental. Hi-C (High-throughput Chromosome Conformation Capture) proximity ligation has emerged as a superior method for direct, high-throughput host identification, overcoming the critical limitations of traditional approaches.
Limitations of Traditional Methods:
- Culture-Based Methods: Require the host to be culturable, missing the estimated >99% of environmental and gut bacteria that are unculturable. They are low-throughput and labor-intensive.
- Indirect Methods (e.g., bioinformatics, CRISPR spacer analysis): Provide only predictive, correlative evidence without physical proof of interaction. They suffer from high false-positive rates and cannot detect active infections in complex samples.
Hi-C Proximity Ligation Mechanism: Hi-C crosslinks physically interacting DNA molecules, including phage DNA within a host bacterium. A proximity ligation step creates chimeric molecules linking phage and host genomes. High-throughput sequencing of these chimeric reads provides direct, physical evidence of phage-host pairs within a natural, complex community, without the need for cultivation.
Quantitative Performance Comparison:
Table 1: Comparison of Phage-Host Linking Methodologies
| Method | Principle | Throughput | Cultivation Required | Direct Physical Link | Key Limitation |
|---|---|---|---|---|---|
| Plaque Assay / Culture | Lysis of bacterial lawn | Very Low | Yes | No | Misses unculturable hosts; low-throughput. |
| Metagenomic Mining | Sequence homology (e.g., tRNA, CRISPR) | High | No | No | Predictive only; high false-positive rate. |
| viralFISH | Fluorescent in situ hybridization | Low | No | Yes (visual) | Low-throughput; difficult in dense samples. |
| Hi-C Proximity Ligation | In situ crosslinking & ligation | Very High | No | Yes (sequenceable) | Requires sufficient co-DNA for ligation. |
Table 2: Representative Hi-C Host-Linking Performance Data
| Study (Sample Type) | Hi-C Protocol | Total Phage-Host Links Identified | % Links to Previously Uncultured Hosts | Key Advantage Demonstrated |
|---|---|---|---|---|
| Gut Microbiome (Human Fecal) | ProxiMeta (Phase Genomics) | 1,824 links | >70% | Uncovered extensive phage-host network in a complex community. |
| Activated Sludge | Hi-C for viral hosts | 148 viral population-host links | ~50% | Linked hosts to novel, non-tailed phages beyond Caudovirales. |
| Marine Virome | MetaHi-C | 352 links | Not specified | Connected hosts to incomplete viral genomes from metagenomes. |
Experimental Protocols
Protocol: Hi-C for Phage-Host Linking from Complex Microbial Communities
I. Sample Fixation and Crosslinking
- Material: Fresh or frozen environmental sample (e.g., fecal material, soil slurry, water filtrate).
- Add formaldehyde to a final concentration of 1-3% and incubate at room temperature for 10-30 minutes with gentle agitation. This crosslinks phage DNA to host DNA inside infected cells.
- Quench crosslinking by adding glycine to a final concentration of 0.125-0.25 M. Incubate for 5-15 minutes at room temperature.
II. Cell Lysis and Chromatin Digestion
- Pellet cells and wash to remove residual formaldehyde.
- Resuspend pellet in appropriate lysis buffer (e.g., containing detergent and protease inhibitors). Incubate to complete lysis.
- Digest crosslinked DNA with a frequent-cutter restriction enzyme (e.g., MboI, HindIII, or Sau3AI) suitable for the expected host genomes.
III. Proximity Ligation and DNA Purification
- Critical Step: Fill in restriction fragment ends with biotinylated nucleotides (e.g., Biotin-14-dATP) using Klenow fragment.
- Perform intra- and inter-molecular ligation under dilute conditions using T4 DNA ligase to favor ligation events between crosslinked fragments. This creates chimeric phage-host DNA molecules.
- Reverse crosslinks by incubating with Proteinase K at 65°C overnight.
- Purify DNA via standard phenol-chloroform extraction and ethanol precipitation.
IV. Biotin Pull-Down and Library Prep
- Shear DNA to ~300-500 bp fragments using a sonicator.
- Capture biotin-labeled chimeric fragments using streptavidin-coated magnetic beads.
- Perform on-bead library preparation for Illumina sequencing, including end repair, adapter ligation, and PCR amplification.
V. Bioinformatics Analysis
- Sequence Processing: Trim adapters, quality filter reads.
- Read Mapping: Map reads to a combined database of curated viral and bacterial genomes/contigs using tools like Bowtie2.
- Link Identification: Identify chimeric reads where one end maps to a viral sequence and the other to a bacterial sequence. Use statistical thresholds (e.g., via tools like
hicstuff,pairsamtools) to filter noise and assign confident phage-host links.
Visualizations
Title: Hi-C Workflow for Phage-Host Linking
Title: Method Comparison Logic
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Reagents for Hi-C Phage-Host Linking
| Item | Function in Protocol | Key Consideration |
|---|---|---|
| Formaldehyde (1-3%) | Crosslinking agent that fixes phage-host DNA physical proximity inside cells. | Concentration and time optimization is critical for efficient crosslinking without over-fixing. |
| Biotin-14-dATP/dCTP | Biotin-labeled nucleotide used to fill in sticky ends after digestion. Labels chimeric molecules for capture. | Essential for selective enrichment of ligation junctions; purity is key. |
| Streptavidin Magnetic Beads | Solid-phase capture of biotin-labeled chimeric DNA fragments. | High binding capacity and low non-specific binding beads improve yield. |
| Frequent-Cutter Restriction Enzyme (e.g., Sau3AI) | Digests crosslinked DNA to create ends for ligation. | Choice influences resolution and bias; should be frequent in host genomes. |
| T4 DNA Ligase | Catalyzes the proximity ligation step, joining crosslinked fragments. | High-concentration, rapid ligase is preferred for efficient chimeric molecule formation. |
| Crosslink Reversal Buffer (w/ Proteinase K) | Reverses formaldehyde crosslinks to release pure DNA for sequencing. | Must include sufficient Proteinase K and incubation time for complete reversal. |
| Reference Database (Viral/Bacterial Genomes) | Curated genome collection for mapping sequencing reads to identify hosts and phages. | Comprehensiveness directly limits discovery; use integrated DBs like RefSeq, GVD, or sample-specific MAGs. |
Application Notes
Hi-C proximity ligation is a revolutionary technique for linking bacteriophages to their bacterial hosts by capturing physical interactions within mixed microbial communities. The interpretation of Hi-C data hinges on understanding key biological and methodological concepts. This note contextualizes these terms within phage-host research.
- Prophages: These are integrated viral genomes within a bacterial chromosome. In Hi-C data, prophages are identified by consistent, high-frequency interaction contacts between phage and host chromosomal DNA, visualized as a dense interaction block off the host's main diagonal. Mapping these links allows for the confident assignment of temperate phages to their specific host strains, even in complex samples.
- Virions: These are the extracellular, virus particles. During Hi-C library preparation, crosslinking captures physical interactions between a virion's packaged DNA and the DNA of the host cell it is attached to or infected. Chimeric reads spanning phage and host sequences from virion-host interactions are critical for identifying lytic or chronic infection cycles.
- Crosslinking (Formaldehyde): The foundational step that freezes in vivo chromatin and phage-host DNA interactions in space and time. Efficient crosslinking is critical for capturing transient virion attachment events and stable prophage integrations, creating the molecular "glue" for subsequent proximity ligation.
- Chimeric Reads: The primary data output of Hi-C. These are sequencing reads containing junctions created by ligating DNA fragments that were spatially proximate. In phage-host linking, a chimeric read that aligns partly to a phage genome and partly to a bacterial genome provides direct evidence of an in situ physical association.
Table 1: Quantitative Signatures of Phage-Host Interactions in Hi-C Data
| Interaction Type | Hi-C Signal Characteristic | Typical Quantitative Metric (from Contact Maps) | Biological Interpretation |
|---|---|---|---|
| Active Prophage | Dense, localized block of interactions off the host diagonal. | Interaction frequency 10-100x higher than background noise in the specific region. | Temperate phage integrated into a specific host chromosome locus. |
| Virion Attachment | Sparse, diffuse network of interactions between phage and host genomic loci. | 1-10 unique chimeric reads linking phage to a specific host; not localized to one chromosomal site. | Virion particle physically attached to cell surface, crosslinked at infection moment. |
| Background Noise | Random, scattered interactions across all genomes. | <1 interaction expected per genomic locus pair after normalization. | Experimental artifact or statistical noise from random ligation. |
Experimental Protocols
Protocol 1: Hi-C Library Preparation for Phage-Host Linking from Environmental Samples
Objective: To capture and sequence crosslinked DNA complexes from a mixed microbial community for subsequent identification of phage-host interactions.
Materials:
- Research Reagent Solutions:
- Fresh 16% Formaldehyde (Methanol-free): For efficient in situ crosslinking.
- Hi-C Ligation Buffer (10X): Contains ATP and Co-factors for efficient blunt-end ligation.
- Biotinylated Pull-down Beads (Streptavidin-coated): For enrichment of chimeric fragments containing a ligation junction.
- Crosslink Reversal Buffer: Proteinase K in EDTA/SDS for digesting proteins and freeing crosslinked DNA.
- HindIII or MluCI (4-cutter) Restriction Enzyme: For chromatin fragmentation, chosen based on host genome frequency.
Method:
- Crosslinking: Concentrate 10-50 ml of environmental sample (e.g., seawater, gut content) by gentle filtration. Resuspend pellet in 1ml PBS. Add 27µl of 16% formaldehyde (final ~1%). Incubate 30 min at room temperature with gentle rotation. Quench with 125µl of 2.5M glycine for 5 min.
- Cell Lysis & Chromatin Digestion: Pellet cells, wash, and lyse using a lysozyme/SDS-based lysis buffer. Use the chosen restriction enzyme (e.g., 100U MluCI) to digest DNA overnight at 37°C.
- Proximity Ligation: Dilute digested lysate in 1X ligation buffer. Perform blunt-end ligation using high-concentration T4 DNA Ligase (100U) for 4 hours at 16°C.
- DNA Purification & Shearing: Reverse crosslinks overnight at 65°C with Proteinase K. Purify DNA via phenol-chloroform extraction. Shear DNA to ~500 bp using a focused-ultrasonicator.
- Biotin Pull-down & Library Prep: Use streptavidin beads to capture biotin-labeled ligation junctions. Prepare sequencing library (end-repair, A-tailing, adapter ligation) on-bead. Elute final library for PCR amplification and sequencing.
Protocol 2:In silicoIdentification of Phage-Host Chimeric Reads
Objective: To bioinformatically process Hi-C sequencing data and extract high-confidence chimeric reads linking phage and host genomes.
Method:
- Preprocessing & Alignment: Trim adapters and low-quality bases. Perform iterative alignment: First, map all reads to a curated phage genome database. Reads that do not map are then aligned to a bacterial/genome database. Use alignment tools (Bowtie2, BWA) with sensitive settings.
- Chimeric Read Extraction: Parse alignment files to identify reads where one segment aligns to a phage contig and the other segment aligns to a bacterial contig with a minimum mapping quality (MAPQ > 20). Discard reads mapping to known common ligation artifacts.
- Interaction Scoring & Visualization: Count unique chimeric read pairs connecting each phage contig to each bacterial contig. Normalize counts by contig length and sequencing depth. Generate an interaction matrix and visualize as a contact map using tools like HiC-Pro or cooler.
Diagrams
Hi-C Phage-Host Linking Workflow
Sources of Phage-Host Chimeric Reads
The Scientist's Toolkit
Table 2: Essential Research Reagents for Hi-C Phage-Host Linking
| Reagent / Material | Function in Protocol |
|---|---|
| Methanol-free Formaldehyde | Ensures efficient in situ crosslinking of DNA-protein and DNA-DNA complexes without shearing. |
| 4- or 6-Cutter Restriction Enzyme (e.g., MluCI, HindIII) | Fragments chromatin at high frequency to increase resolution and likelihood of capturing phage-host junctions. |
| T4 DNA Ligase (High-Concentration) | Catalyzes the blunt-end ligation of crosslinked, digested DNA fragments in dilute conditions to favor proximity ligation. |
| Biotin-14-dATP | Incorporated during fill-in of restriction overhangs, labeling the ligation junction for streptavidin-based enrichment. |
| Streptavidin-coated Magnetic Beads | Selectively captures biotinylated chimeric fragments, reducing background non-ligated DNA for cleaner libraries. |
| Phage & Host Genome Databases | Curated, comprehensive sequence databases for iterative read alignment to identify chimeric pairs. |
| Crosslink Reversal Buffer (Prot. K/SDS) | Digests crosslinking proteins and reverses formaldehyde adducts to release pure DNA for downstream processing. |
Step-by-Step Protocol: Implementing Hi-C to Uncover Phage Hosts in Complex Samples
Sample Preparation Strategies for Environmental, Clinical, and Synthetic Communities
This document details sample preparation strategies for complex microbial communities, framed within the overarching thesis of applying Hi-C proximity ligation to elucidate phage-host interactions. The accurate linking of bacteriophages to their bacterial hosts is critical for understanding microbial ecology, phage therapy development, and antimicrobial discovery. Hi-C methodology, which cross-links physically interacting DNA strands in situ, provides a powerful tool for this linkage but is profoundly dependent on the initial sample preparation to preserve native interactions and yield high-quality, representative DNA.
The optimal preparation strategy varies significantly by community origin. The primary goal across all types is to stabilize intimate phage-bacteriome contacts while minimizing exogenous contamination and bias.
Table 1: Strategic Comparison by Community Type
| Community Type | Primary Challenge | Key Preparation Focus | Optimal Stabilization Method |
|---|---|---|---|
| Environmental (e.g., soil, seawater) | Inhibitory substances (humics, salts), low biomass | Efficient cell collection & inhibitor removal | In-situ crosslinking with formaldehyde followed by filtration or centrifugation. |
| Clinical (e.g., sputum, stool) | Host human DNA contamination, ethical/biosafety constraints | Depletion of host cells/DNA, pathogen inactivation | Density gradient centrifugation, selective lysis, or use of commercial host depletion kits prior to crosslinking. |
| Synthetic (Defined co-cultures) | Precise control of interaction timing & ratios | Synchronization of infection cycles | Controlled crosslinking at specific Multiplicity of Infection (MOI) and time post-infection in bioreactors. |
Detailed Application Notes and Protocols
Protocol 3.1: Environmental Water Sample Preparation for Hi-C
Application Note: Designed for aquatic environments (lakes, wastewater) to capture native phage-host complexes.
Materials & Reagents:
- Sterile Filtration Unit (0.22 µm pore, followed by 0.1 µm or 100 kDa tangential flow filter): Sequential size-based collection of bacterial cells and associated phages.
- Crosslinking Solution: 3% Formaldehyde (v/v) in filtered site water (pH ~7.0).
- Glycine Quench Solution: 1.25 M glycine (sterile).
- PBS-Mg Buffer: 1X PBS with 10 mM MgCl₂ (maintains capsid integrity).
Procedure:
- In-situ Crosslinking: Immediately upon collection, add crosslinking solution to the water sample to a final concentration of 1% formaldehyde. Incubate at room temperature for 30 minutes with gentle agitation.
- Quenching: Add glycine to a final concentration of 125 mM to quench crosslinking. Incubate 10 min at RT.
- Biomass Concentration: Pass the quenched sample through a 0.22 µm filter to capture bacterial cells. Subsequently, pass the filtrate through a 100 kDa filter to concentrate phage particles.
- Combination: Resuspend both filters in 5 mL of ice-cold PBS-Mg buffer, combining the fractions.
- Storage: Pellet combined biomass at 8,000 x g, 10 min, 4°C. Flash-freeze pellet in liquid N₂ and store at -80°C until Hi-C library construction.
Protocol 3.2: Clinical Stool Sample Preparation with Host Depletion
Application Note: Focuses on human gut microbiome, prioritizing biosafety and reducing human DNA background >90%.
Materials & Reagents:
- Anaerobic Transport Medium: For preserving anaerobic taxa during transport.
- Host Cell Depletion Kit: e.g., MICROBEnrich or HostZERO.
- Inactivation/Stabilization Buffer: e.g., DNA/RNA Shield with 2% formaldehyde.
- Differential Centrifugation Buffers: Sucrose gradient buffers (10%-40%).
Procedure:
- Inactivation & Stabilization: Homogenize 1g stool in 10 mL inactivation/stabilization buffer. Incubate 1 hour at 4°C.
- Coarse Removal: Centrifuge at 500 x g for 5 min to pellet large debris and eukaryotic cells. Transfer supernatant.
- Microbial Enrichment: Use a commercial host depletion kit per manufacturer's instructions, OR perform density gradient centrifugation: layer supernatant on a 10%-40% sucrose gradient, centrifuge at 2,000 x g for 20 min. Harvest the interphase layer containing microbial cells.
- Wash & Crosslink: Pellet enriched microbial cells at 8,000 x g for 10 min. Wash twice with PBS. Resuspend in PBS with 1% formaldehyde for final crosslinking (20 min, RT). Quench with glycine.
- Pellet & Store: Pellet cells, wash, flash-freeze, and store at -80°C.
Protocol 3.3: Synthetic Community Hi-C Sample Preparation
Application Note: For defined phage-bacteria co-cultures, enabling precise study of infection dynamics.
Materials & Reagents:
- Bioreactor or Controlled Environment Chamber: For precise growth control.
- Synchronization Agents: e.g., Mitomycin C for prophage induction.
- Crosslinking Agent: 3% Formaldehyde in growth medium.
- Stop Solution: 125 mM Glycine in PBS.
Procedure:
- Culture Synchronization: Grow bacterial host to mid-log phase (OD₆₀₀ ~0.3-0.4). Add phage at a defined MOI (e.g., MOI=5) or inducing agent.
- Infectious Interaction: Allow adsorption for 15-30 min. Add fresh medium to dilute unadsorbed phage. Incubate for a predetermined time-post-infection (e.g., 25 min for early interactions).
- Precise Crosslinking: Rapidly add formaldehyde to culture to 1% final concentration. Incubate exactly 10 min at RT with shaking.
- Quenching & Harvest: Add glycine to 125 mM final concentration. Incubate 5 min. Centrifuge culture at 8,000 x g for 10 min at 4°C.
- Wash & Store: Wash pellet twice with ice-cold PBS. Flash-freeze and store at -80°C.
The Scientist's Toolkit: Key Reagent Solutions
Table 2: Essential Research Reagents for Hi-C Sample Prep
| Reagent / Solution | Function in Hi-C Prep | Key Consideration |
|---|---|---|
| Formaldehyde (1-3%) | Crosslinking agent that creates covalent bonds between proximal DNA strands inside cells, freezing phage-host contacts. | Concentration and time are critical; over-fixing reduces DNA yield and accessibility. |
| Glycine (125 mM) | Quenches formaldehyde by reacting with excess reagent, stopping the crosslinking process. | Essential for reproducible and controllable fixation. |
| DNA/RNA Shield (Zymo) | Inactivates nucleases and pathogens while stabilizing nucleic acids. Useful for hazardous clinical samples. | Allows safe handling without immediate freezing. |
| Host Depletion Kits (e.g., MICROBEnrich) | Selectively lyses human eukaryotic cells or binds human DNA, enriching for microbial and viral biomass. | Critical for increasing sequencing depth on target communities in clinical samples. |
| Sucrose or Nycodenz Gradients | Separates microbial cells from denser eukaryotic debris and less dense vesicles/virions via density. | A physical method for host depletion, complementary to kits. |
| PBS with MgCl₂ (10mM) | Wash and resuspension buffer that helps maintain the integrity of phage capsids and bacterial membranes. | Prevents premature lysis and loss of phage DNA. |
Visualized Workflows and Pathways
Title: Hi-C Sample Prep: Environmental Water
Title: Clinical Stool Prep with Host Depletion
Title: Synthetic Community Infection & Crosslinking Timeline
Application Notes
Within the broader thesis employing Hi-C proximity ligation to map phage-host interaction networks, the in situ formaldehyde crosslinking step is foundational. It captures transient, physical contacts between the infecting phage DNA and the host bacterial chromosome at a specific moment in the infection cycle. This covalent "freezing" preserves the three-dimensional proximity architecture for downstream processing, enabling the identification of host genomic loci that are spatially adjacent to the phage genome. The efficiency and specificity of this crosslinking directly determine the signal-to-noise ratio in the final contact maps, making optimization critical for distinguishing true integration or interaction sites from random ligation events.
Key Quantitative Parameters for Crosslinking Optimization
The following tables summarize critical data from recent literature on optimizing formaldehyde crosslinking for chromatin interaction studies in prokaryotes, adapted for phage-host systems.
Table 1: Formaldehyde Crosslinking Parameters and Outcomes
| Parameter | Typical Range | Optimal Value for Prokaryotic Hi-C | Effect on Results |
|---|---|---|---|
| Formaldehyde Concentration | 0.5% - 3% | 1% - 2% | Higher conc. increases crosslink yield but can reduce ligation efficiency. |
| Crosslinking Temperature | 4°C - 37°C | Room Temp (20-25°C) | Balances reaction kinetics with preservation of native state. |
| Crosslinking Duration | 5 min - 30 min | 10 - 20 min | Shorter times may under-crosslink; longer times can over-crosslink. |
| Quenching Agent | Glycine, Tris | 125 mM Glycine | Stops reaction, prevents protein-nucleic acid over-crosslinking. |
| Cell Density (OD600) | 0.2 - 1.0 | 0.4 - 0.6 | Ensures even crosslinking and avoids cell clumping. |
Table 2: Impact of Crosslinking on Downstream Hi-C Metrics
| Metric | Under-crosslinked Sample | Optimally-crosslinked Sample | Over-crosslinked Sample |
|---|---|---|---|
| Ligation Efficiency | High but non-specific | High and specific | Very Low |
| Valid Read Pairs | Low percentage (< 10%) | High percentage (20-40%) | Extremely Low |
| Signal-to-Noise (Trans/Cis ratio) | Low (< 0.1) | High (> 0.5) | Not detectable |
| Peak Sharpness at Interaction Loci | Broad, diffuse | Sharp, defined | No peaks |
Experimental Protocols
Protocol 1:In situFormaldehyde Crosslinking of Phage-Infected Bacterial Cultures
Objective: To covalently fix phage-host genomic contacts within infected bacterial cells prior to Hi-C library preparation.
Materials:
- Phage-infected bacterial culture at desired post-infection time point.
- 16% Formaldehyde Solution (methanol-free, molecular biology grade).
- 2.5M Glycine (sterile filtered).
- PBS or appropriate cold buffer (e.g., 10 mM Tris-HCl pH 8.0, 100 mM NaCl).
- Ice-cold water bath.
- Centrifuge and rotors for bacterial cell pelleting.
Procedure:
- Culture Preparation: Grow the bacterial host to mid-log phase (OD600 ~0.3-0.4). Infect with phage at a defined multiplicity of infection (MOI). Incubate under appropriate conditions until the target time point post-infection (e.g., early, mid, or late infection).
- Crosslinking Initiation: For 10 ml of infected culture, directly add 16% formaldehyde to a final concentration of 1% (e.g., 625 µl). Mix immediately and thoroughly by inversion or gentle vortexing.
- Incubation: Incubate the reaction at room temperature for 15 minutes with gentle rotation or occasional shaking to keep cells suspended.
- Quenching: Add 2.5M glycine to a final concentration of 0.125 M (e.g., 500 µl per 10 ml). Mix thoroughly. Incubate at room temperature for 5 minutes to quench unreacted formaldehyde.
- Cell Harvesting: Transfer the crosslinked culture to a centrifuge tube on ice. Pellet cells at 4,000 x g for 10 minutes at 4°C. Carefully decant the supernatant.
- Washing: Resuspend the cell pellet in 10 ml of ice-cold PBS (or cold Tris-NaCl buffer). Repeat centrifugation. Perform two total washes to ensure complete removal of glycine and formaldehyde.
- Storage: After the final wash, flash-freeze the cell pellet in liquid nitrogen or a dry-ice ethanol bath. Store at -80°C until ready for Hi-C library preparation.
Protocol 2: Integration into Hi-C Workflow: Cell Lysis and Chromatin Fragmentation
Objective: To process crosslinked cells for proximity ligation, beginning with lysis and fragmentation of crosslinked chromatin.
Materials:
- Crosslinked cell pellet (from Protocol 1).
- Lysis Buffer: 10 mM Tris-HCl pH 8.0, 50 mM NaCl, 1x EDTA-free protease inhibitor cocktail, 0.5% SDS.
- 20% Triton X-100.
- Restriction enzyme with frequent recognition site in host and phage genomes (e.g., MluCI, HinP1I) and corresponding 10x buffer.
- Water bath or thermal mixer.
- Ice.
Procedure:
- Cell Lysis: Thaw the crosslinked pellet on ice. Resuspend completely in 1 ml of cold Lysis Buffer. Incubate for 30 minutes at 37°C with gentle agitation to lyse cells and solubilize crosslinked nucleoprotein complexes.
- SDS Quenching: Add 55 µl of 20% Triton X-100 (to a final concentration of ~1%). Mix thoroughly. Incubate for 1 hour at 37°C with agitation to sequester SDS, which would otherwise inhibit the subsequent restriction enzyme.
- Chromatin Digestion: Distribute the lysate into aliquots for digestion. Add 10x restriction enzyme buffer and 50-100 units of restriction enzyme per aliquot. Incubate overnight at 37°C with gentle rotation.
- Enzyme Inactivation: The following day, incubate the digest at 65°C for 20 minutes to inactivate the restriction enzyme. Proceed immediately to end-repair, biotinylation, and ligation steps as per standard Hi-C protocols (e.g., using a commercial Hi-C kit or published in-house methods).
Diagrams
Title: Workflow: From Phage Infection to Hi-C Contact Map
Title: Chemistry of FA Crosslinking Phage-Host Contacts
The Scientist's Toolkit
Table 3: Essential Research Reagent Solutions for Phage-Host Hi-C
| Reagent / Material | Function / Role in Protocol | Key Considerations |
|---|---|---|
| Methanol-free Formaldehyde (16%) | The crosslinking agent. Creates methylene bridges between primary amines in proteins and nucleic acids. | Methanol-free grade prevents protein precipitation and non-specific crosslinking. Aliquot and store at -20°C. |
| Glycine (2.5M stock) | Quenching agent. Terminates crosslinking by reacting with excess formaldehyde. | Must be sterile-filtered. Critical for preventing over-crosslinking, which inhibits digestion/ligation. |
| Frequent-Cutter Restriction Enzyme (e.g., HinP1I) | Fragments crosslinked chromatin for proximity ligation. Creates cohesive ends. | Choose an enzyme with high frequency in both host and phage genomes (4-6 bp cutter). Verify activity in Triton X-100 buffer. |
| Triton X-100 (20% solution) | Non-ionic detergent used to quench SDS after lysis, enabling restriction enzyme activity. | Ensures complete sequestration of SDS from the lysis step. |
| Biotin-14-dATP/dCTP | Labels fragment ends during fill-in for selective pull-down of ligated junctions. | Essential for enriching for chimeric fragments representing cross-ligated phage-host contacts. |
| Streptavidin Magnetic Beads | Captures biotinylated ligation junctions post-ligation for library construction. | High binding capacity and low non-specific DNA binding are crucial for yield and purity. |
| Phase-lock Gel Tubes | Facilitates clean phenol:chloroform extractions of crosslinked DNA/protein. | Particularly useful during the initial lysate cleanup steps to recover fragile crosslinked complexes. |
Enzymatic Digestion, Proximity Ligation, and DNA Purification Workflow
The identification of bacteriophage-host interaction networks is critical for understanding microbial ecology and developing phage-based therapies. Hi-C proximity ligation, adapted for phage-host research, enables the detection of physical interactions between phage and bacterial genomic DNA within infected cells. This workflow captures chromosomal conformation data, revealing which bacterial hosts specific phages are infecting in complex communities. The protocol detailed herein is designed for the rigorous preparation of proximity-ligated DNA libraries suitable for high-throughput sequencing and subsequent bioinformatic linking of phages to their hosts.
Research Reagent Solutions: Essential Materials
The following table lists key reagents and their specific functions in the Hi-C protocol for phage-host linking.
| Reagent / Material | Function in the Workflow |
|---|---|
| Formaldehyde (2-3%) | Crosslinking agent that fixes phage-host DNA complexes in spatial proximity. |
| SDS (Sodium Dodecyl Sulfate) | Ionic detergent for cell lysis and denaturation of proteins post-crosslinking. |
| DpnII / MluCI / HindIII | Restriction enzymes (frequent cutters) for digesting crosslinked DNA into fragments. |
| Biotin-14-dATP | Labeling nucleotide incorporated into digested DNA ends to mark ligation junctions. |
| T4 DNA Ligase | Enzyme facilitating intra-molecular ligation of crosslink-stabilized, digested DNA ends. |
| Streptavidin-coated Magnetic Beads | Solid-phase support for purification of biotin-labeled ligation junctions. |
| Proteinase K | Protease for reversing formaldehyde crosslinks by digesting proteins. |
| AMPure XP or SPRI Beads | Magnetic beads for size selection and purification of DNA libraries. |
| Phusion High-Fidelity DNA Polymerase | PCR amplification of purified ligation products for sequencing library construction. |
| DynaMag-2 Magnet | Magnetic rack for separations involving magnetic beads. |
Detailed Experimental Protocol
Cell Culture, Infection, and Crosslinking
- Grow the bacterial culture of interest to mid-log phase (OD600 ~0.4-0.6).
- Infect with phage at a desired Multiplicity of Infection (MOI, e.g., 1-10). Incubate for a specific time post-infection (e.g., 15-30 mins).
- Add fresh, chilled formaldehyde to a final concentration of 2% (v/v) directly to the culture. Mix well.
- Incubate at room temperature for 20-30 minutes with gentle rotation to crosslink DNA-protein and DNA-DNA complexes.
- Quench the crosslinking reaction by adding glycine to a final concentration of 0.2 M. Incubate for 5-10 minutes at room temperature.
- Pellet cells by centrifugation (4,000 x g, 10 min, 4°C). Wash cell pellet twice with 1x cold PBS.
Cell Lysis and DNA Digestion
- Resuspend the cell pellet in Lysis Buffer (50 mM Tris-HCl pH 8.0, 50 mM NaCl, 1% SDS, protease inhibitors). Incubate for 15-30 minutes at 37°C.
- Dilute the SDS concentration to ~0.1% by adding 1x NEBuffer appropriate for the chosen restriction enzyme and water.
- Add 1% (v/v) Triton X-100 to sequester SDS. Mix thoroughly.
- Add the chosen frequent-cutter restriction enzyme (e.g., 200-400 units of DpnII). Incubate overnight (16-18 hours) at 37°C with gentle agitation.
Proximity Ligation
- Inactivate the restriction enzyme by incubating at 65°C for 20 minutes.
- Cool the reaction to room temperature. Prepare a master mix containing: 1x T4 DNA Ligase Buffer, 1 mM ATP, 1% Triton X-100, 0.1 mg/mL BSA, and a low concentration of biotin-14-dATP.
- Add the master mix and a high concentration of T4 DNA Ligase (e.g., 100 Weiss units) to the digested DNA. Mix gently.
- Perform ligation at room temperature for 4-6 hours or overnight at 16°C. This promotes intra-molecular ligation of crosslinked ends.
DNA Purification & Biotin Pull-Down
- Reverse crosslinks by adding Proteinase K to a final concentration of 0.2 mg/mL and SDS to 0.5%. Incubate at 55°C for 30 minutes, then at 68°C overnight.
- Purify DNA using a standard phenol:chloroform:isoamyl alcohol extraction and ethanol precipitation. Resuspend in TE buffer.
- Shear the purified DNA to an average fragment size of 300-500 bp using a focused-ultrasonicator (Covaris).
- Perform an end-repair and A-tailing reaction on the sheared DNA using standard kits.
- Bind the DNA to pre-washed Streptavidin-coated magnetic beads in high-salt buffer (1 M NaCl) for 15 minutes at room temperature.
- Wash beads twice with 1x B&W Buffer (5 mM Tris-HCl pH 7.5, 0.5 mM EDTA, 1 M NaCl, 0.1% Tween-20) and once with 1x Low Salt TE (10 mM Tris-HCl, 0.1 mM EDTA).
- Elute non-biotinylated fragments in low salt buffer. The biotinylated ligation junctions remain bound.
Library Preparation for Sequencing
- On-bead, ligate Illumina sequencing adapters to the bound DNA fragments using T4 DNA Ligase.
- Perform a final wash to remove excess adapters.
- Amplify the library directly on the beads via PCR using Phusion High-Fidelity DNA Polymerase and indexed primers.
- Purify the final library using AMPure XP beads for size selection (e.g., 0.8x ratio to remove large fragments, then 1.2x ratio to recover the target library).
- Quantify the library using Qubit and assess size distribution with a Bioanalyzer/TapeStation.
- Sequence on an Illumina platform (typically paired-end 150 bp).
Table 1: Typical Yield Metrics at Critical Protocol Steps
| Protocol Step | Typical DNA Yield (from 10^8 E. coli cells) | Notes / Quality Check |
|---|---|---|
| Post-Crosslinking & Lysis | 5-10 µg | Assessed by Nanodrop; A260/A280 ~1.8. |
| Post-Restriction Digestion | 4-9 µg | Run on gel to check smear; reduced viscosity. |
| Post-Proximity Ligation & De-crosslinking | 3-7 µg | |
| Post-Shearing & Size Selection | 1-2 µg | Bioanalyzer profile: peak ~350 bp. |
| Final PCR-Amplified Library | 50-200 ng | Ready for sequencing; must pass Bioanalyzer QC. |
Table 2: Key Reaction Conditions and Parameters
| Reaction | Key Components | Incubation Conditions | Duration |
|---|---|---|---|
| Crosslinking | 2% Formaldehyde, Culture Media | Room Temp, Rotation | 20-30 min |
| Restriction Digest | DpnII (400 U), 1x NEBuffer, Triton X-100 | 37°C, Gentle Agitation | 16-18 hrs |
| Proximity Ligation | T4 DNA Ligase (100 U), biotin-14-dATP, Ligase Buffer | 16°C or Room Temp | 4-6 hrs or O/N |
| Crosslink Reversal | Proteinase K (0.2 mg/mL), 0.5% SDS | 55°C, then 68°C | 30 min, then O/N |
| Adapter Ligation (On-bead) | Illumina Adapters, T4 DNA Ligase | 20°C | 15 min |
| Library Amplification | Phusion Polymerase, Indexed Primers | 98°C/10s, 65°C/30s, 72°C/30s | 12-15 cycles |
Workflow and Pathway Diagrams
Diagram 1: Hi-C for Phage-Host Linking Workflow
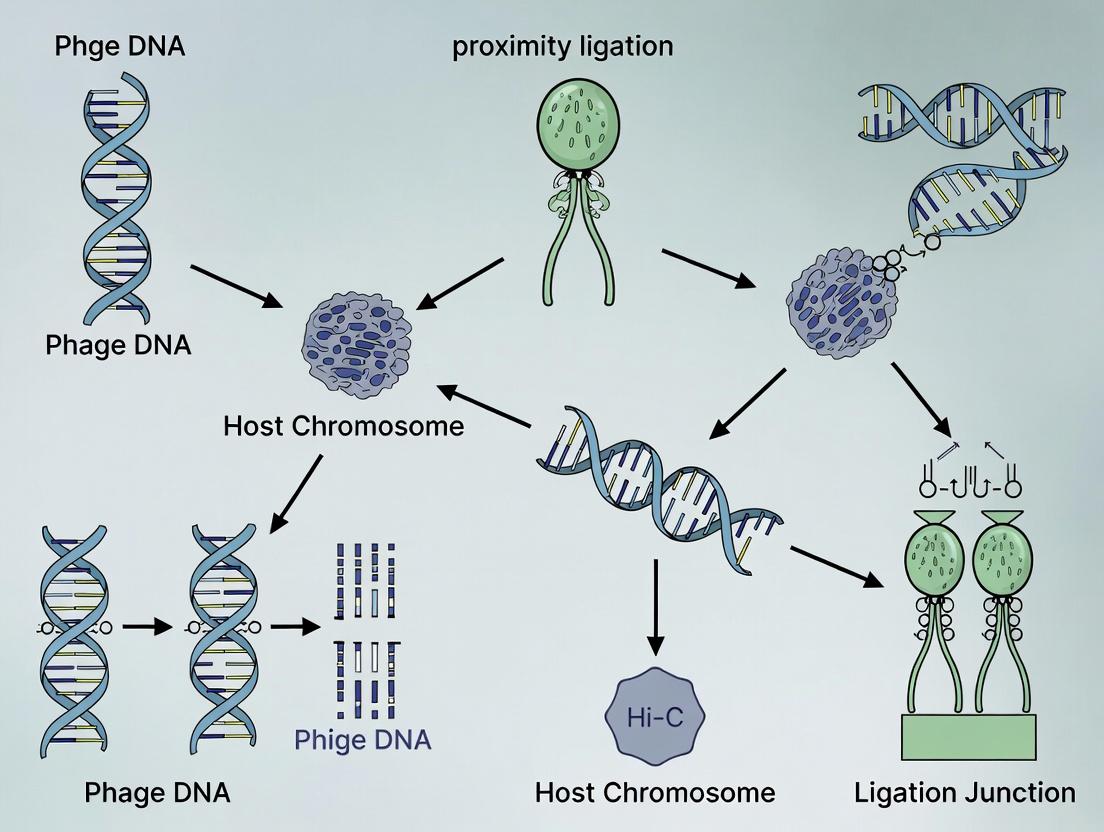
Diagram 2: Molecular Basis of Phage-Host Linking via Hi-C
1. Introduction Within a thesis on Hi-C proximity ligation for phage host linking, optimizing sequencing parameters is critical for deconvoluting complex microbial communities and confidently linking phages to their bacterial hosts. This application note details the considerations for sequencing depth, read length, and library construction protocols to ensure high-resolution, statistically robust data for downstream network analysis and therapeutic discovery.
2. Key Considerations & Quantitative Summary
Table 1: Sequencing Parameter Guidelines for Phage-Host Hi-C
| Parameter | Recommended Specification | Rationale for Phage-Host Linking |
|---|---|---|
| Sequencing Depth | 50-100 million paired-end reads per sample (complex community) | Ensures sufficient coverage of low-abundance phage-host interactions; statistical power for linking. |
| Read Length | 2 x 150 bp (PE150) minimum; 2 x 250 bp preferred. | Long reads aid in spanning repetitive regions and improving alignment specificity of chimeric reads. |
| Library Insert Size | 300-500 bp. | Optimizes capture of cross-linked DNA fragments while maintaining efficient cluster generation on flow cells. |
| Sequencing Type | Paired-end (PE), Illumina platform. | Provides sequence from both ends of the insert, crucial for mapping chimeric junctions. |
| Read Type | Must include non-duplicate, properly paired, and chimeric reads. | Chimeric reads are the direct evidence of proximity ligation events. |
Table 2: Impact of Parameters on Data Output
| Parameter | Insufficient/Suboptimal | Optimal | Excessive |
|---|---|---|---|
| Depth | Missed rare links, low statistical confidence. | Robust interaction detection, saturation of significant contacts. | Diminishing returns, increased cost. |
| Read Length | Ambiguous alignments, missed junctions. | Confident alignment of both read ends across junction. | Minimal added value for standard Hi-C. |
| Insert Size | Over-representation of unligated fragments. | Balanced yield of intra- and inter-molecular ligations. | Reduced complexity, potential bias. |
3. Detailed Experimental Protocol: Hi-C Library Construction for Phage-Host Samples
Protocol: In situ Hi-C for Microbial Communities Adapted from Marbouty et al., 2021 and current best practices.
A. Crosslinking and Lysis
- Fixation: Mix environmental sample or co-culture with fresh 3% formaldehyde. Incubate at room temperature for 30 min with gentle rotation.
- Quenching: Add glycine to a final concentration of 0.25 M. Incubate for 15 min at RT.
- Pellet cells: Centrifuge at 4°C. Wash pellet 2x with cold PBS.
- Lysis: Resuspend pellet in cold lysis buffer (50 mM Tris-HCl pH 8.0, 50 mM Sucrose, 100 mM NaCl, 1% Triton X-100, 1x protease inhibitor). Incubate on ice for 30 min.
B. Chromatin Digestion and Marking
- Pellet nuclei/phages: Centrifuge lysate. Resuspend in 1x NEBuffer 3.1.
- Digest: Add 100 U of HindIII (or 4-cutter like DpnII for finer resolution) per sample. Incubate at 37°C overnight with gentle agitation.
- Fill & Mark: Heat inactivate at 65°C for 20 min. Add Biotin-14-dATP and Klenow Fragment (exo-) to fill 5’ overhangs. Incubate at 37°C for 90 min.
C. Proximity Ligation
- Dilute & Ligate: Dilute reaction with ligation buffer to favor inter-molecular ligation. Add T4 DNA Ligase. Incubate at 16°C for 4-6 hours.
- Reverse Crosslinks: Add Proteinase K and SDS. Incubate at 65°C overnight.
- DNA Purification: Perform Phenol:Chloroform:IAA extraction followed by ethanol precipitation. Resuspend in TE buffer.
D. Biotin Capture and Library Prep
- Shearing: Sonicate DNA to ~400 bp average fragment size.
- Size Selection: Perform double-sided SPRI bead cleanup to select 300-500 bp fragments.
- Biotin Pull-down: Incubate with Streptavidin-coated magnetic beads in binding buffer (1 M NaCl, 5 mM Tris-HCl pH 8.0, 0.5 mM EDTA). Wash thoroughly.
- On-Bead Library Prep: Perform end-repair, A-tailing, and adapter ligation directly on beads. Include dual-indexed adapters for multiplexing.
- Final PCR: Perform limited-cycle PCR (8-12 cycles) to amplify the library. Purify with SPRI beads.
- QC: Quantify by Qubit and analyze fragment size distribution by Bioanalyzer/TapeStation.
4. Visualization: Experimental Workflow & Data Analysis Logic
Hi-C Proximity Ligation Experimental Workflow
Bioinformatics Pipeline for Phage-Host Link Identification
5. The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for Phage-Host Hi-C Experiments
| Item | Function & Rationale |
|---|---|
| Formaldehyde (3%) | Crosslinks phage particles to host DNA at the site of infection, capturing physical proximity. |
| HindIII or DpnII (NEB) | Restriction enzymes to digest crosslinked chromatin, defining Hi-C resolution. |
| Biotin-14-dATP | Labels digested DNA ends for subsequent streptavidin-based enrichment of ligation junctions. |
| T4 DNA Ligase (High-Concentration) | Performs intra- and inter-molecular ligation of crosslinked, biotinylated ends under dilute conditions. |
| Streptavidin Magnetic Beads | Captures biotinylated ligation products, removing background non-ligated DNA. |
| Dual-Indexed Adapters (Illumina) | Allows multiplexing of multiple samples in a single sequencing run. |
| SPRIselect Beads | For precise size selection and cleanup during library construction. |
| Phage & Host Genome Databases | Curated reference sequences for accurate dual-alignment of chimeric reads. |
This protocol details a downstream bioinformatics pipeline for processing sequencing data derived from Hi-C proximity ligation experiments. Within the broader thesis on using Hi-C for phage host linking, this pipeline is critical for translating raw sequence data into statistically robust physical contacts between phage and host genomes, enabling the discovery and validation of novel phage-host relationships for therapeutic development.
Pipeline Workflow & Protocols
Diagram 1: Hi-C host linking bioinformatics workflow (78 chars)
Protocol 2.1: Initial Quality Control and Trimming
- Tool:
fastp(version 0.23.4) - Command:
- Purpose: Removes low-quality bases, adapter sequences, and polyG tails. Generates a QC report.
Protocol 2.2: Alignment to Composite Reference Genome
- Tool:
Bowtie2(version 2.5.3) - Reference Preparation: Create a composite FASTA file containing all potential bacterial host genomes and known phage genomes.
Alignment Command:
Post-alignment Processing: Convert SAM to sorted BAM and index.
Protocol 2.3: Hi-C Contact Filtering and Deduplication
- Tool:
pairtools(version 1.0.3) - Workflow:
- Purpose: Isolates bona fide Hi-C contact pairs, removing technical noise.
Protocol 2.4: In-Silico Enrichment for Phage-Host Contacts
- Custom Python Script:
extract_chimeric_pairs.py
- Logic: Parse the
.pairs file to extract read pairs where one read aligns to a phage contig and the other aligns to a bacterial contig.
- Key Output: A table listing all phage-host read pairs with genomic coordinates and alignment scores.
Protocol 2.5: Statistical Host Assignment
- Method: Binomial Test or Hypergeometric Test against background noise.
- Implementation (R):
- Assignment Threshold: Adjusted p-value < 0.05 and contact count > 5.
Data Presentation
Table 1: Key Performance Metrics from a Representative Hi-C Host-Linking Run
Metric
Value
Interpretation
Raw Read Pairs
50,000,000
Total sequencing depth
Post-QC Read Pairs
48,500,000 (97%)
High-quality input data
Aligned Pairs (Composite Ref)
35,150,000 (72.5%)
Efficient alignment
Valid Hi-C Pairs
8,432,000 (24%)
Typical yield for complex metagenome
Phage-Host Chimeric Pairs
12,450
Candidate interactions
Significant Assignments (FDR<0.05)
15 Phage 8 Hosts
High-confidence links
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Materials and Tools for Hi-C Host-Linking Analysis
Item
Function & Rationale
Proximity Ligation Kit (e.g., Arima-HiC)
Standardizes crosslinking, digestion, and biotin fill-in for reproducible contact capture.
Size Selection Beads (SPRI)
Critical for isolating correctly ligated fragments (~300-700 bp) post-digestion.
Biotin Capture Streptavidin Beads
Enriches for fragments containing the biotin-labeled ligation junction.
High-Fidelity PCR Master Mix
Amplifies library post-capture with minimal bias for NGS preparation.
Composite Reference Database
Custom FASTA of all relevant host genomes and phage/virome sequences; essential for alignment.
High-Performance Computing (HPC) Cluster
Necessary for memory-intensive alignment and processing of large metagenomic Hi-C datasets.
Dedicated Bioinformatics Pipeline (Snakemake/Nextflow)
Ensures reproducibility, scalability, and automated execution of the multi-step protocol.
Logical Decision Pathway for Host Assignment
Diagram 2: Decision logic for phage host assignment (81 chars)
*Genomic evidence includes CRISPR spacer matches, tRNA similarity, or sequence homology.
Application Notes
Within the thesis framework of using Hi-C proximity ligation to link bacteriophages (phages) to their bacterial hosts in complex samples, the derived data finds direct, high-impact applications in two critical areas: the rational design of therapeutic phage cocktails and the profiling of antibiotic resistance genes (ARGs) within a functional host context.
1. Application: Rational Phage Cocktail Design Traditional phage isolation and host range determination are low-throughput and often fail to capture the true interaction network in microbial communities. Hi-C phage-host linking provides a snapshot of which phages are actively infecting which bacterial strains in situ. This enables data-driven cocktail design.
- Key Data Points: Hi-C generates quantitative linkage frequencies between phage and host genomes (Table 1). Strong, consistent linkages indicate a robust, active host relationship.
- Rationale: A therapeutic cocktail should target the maximum diversity of pathogenic strains (breadth) while minimizing the number of phages needed (efficiency). By analyzing linkage networks, researchers can select a minimal set of phages whose combined host ranges, as empirically defined by Hi-C links, cover all target pathogen strains present in a sample (e.g., a chronic wound microbiome). This moves beyond in vitro plaque assays to leverage ecological interaction data.
2. Application: Functional Antibiotic Resistance Profiling Metagenomic sequencing can catalog all ARGs in a sample but cannot determine which bacterial hosts carry them, crucial for understanding resistance reservoirs and transmission. Integrating Hi-C host linking with ARG annotation solves this.
- Key Data Points: Hi-C links physically connect ARG-containing DNA fragments to the bacterial genome of origin (Table 2). This allows for the creation of a resistome map tagged to specific hosts.
- Rationale: Identifying which specific bacterial taxa harbor clinically relevant ARGs (e.g., ESBLs, carbapenemases) informs risk assessment and treatment strategies. Furthermore, linking temperate phages to hosts carrying ARGs can identify potential vectors for horizontal gene transfer, profiling the mobile resistome.
Protocols
Protocol 1: Hi-C Proximity Ligation for Phage-Host and ARG Host Linking from Microbial Communities
Title: Sample processing, crosslinking, and proximity ligation to capture phage-host genomic interactions.
Research Reagent Solutions & Essential Materials:
| Item | Function |
|---|---|
| Crosslinking Buffer (3% formaldehyde in 1X PBS) | Fixes physical interactions between phage DNA and host bacterial chromosome inside the cell. |
| Hi-C Ligation Master Mix (T4 DNA Ligase buffer, ATP, T4 DNA Ligase, 10% Triton X-100) | Ligates crosslinked, compatible ends of crosslinked DNA fragments in situ. |
| Biotin-14-dATP | Labels ligation junctions during fill-in for subsequent streptavidin-based pulldown. |
| Streptavidin-coated Magnetic Beads | Isolates biotinylated chimeric fragments containing phage-host ligation products. |
| Phase Lock Gel Tubes | Improves phenol:chloroform separation of crosslinked DNA during extraction. |
| Chromatin Shearing Covaris ultrasonicator | Shears crosslinked DNA to optimal size (~300-500 bp) for sequencing library construction. |
Detailed Methodology:
- Sample Fixation: Concentrate 10^9 - 10^10 microbial cells from environmental or clinical sample (e.g., filtered water, homogenized sputum). Resuspend pellet in 10 ml cold PBS. Add 540 µl of 37% formaldehyde to final 2% v/v. Incubate 30 min at room temperature with gentle rotation.
- Quenching & Wash: Add 1.25 ml of 2.5M glycine to quench. Incubate 5 min. Pellet cells (4000 x g, 5 min, 4°C). Wash pellet 2x with cold PBS.
- Cell Lysis & Chromatin Digestion: Resuspend pellet in 1 ml lysis buffer (10mM Tris-HCl pH8.0, 10mM NaCl, 0.2% Igepal CA-630, protease inhibitors). Lyse on ice for 30 min. Pellet nuclei/protein-DNA complexes (5000 x g, 5 min, 4°C). Resuspend in 0.5 ml 1X NEBuffer 3.1. Add 25 µl 10% SDS, incubate 10 min at 65°C. Add 100 µl 20% Triton X-100 to sequester SDS. Digest chromatin with 100 U MluCI-HF restriction enzyme (or other 4-cutter) overnight at 37°C.
- Fill-in & Proximity Ligation: Fill in restriction overhangs and label junctions by adding 37.5 µl of a mix containing 0.25 mM biotin-14-dATP, 0.25 mM dCTP, 0.25 mM dGTP, 0.25 mM dTTP, and 50 U DNA Polymerase I, Large (Klenow) Fragment. Incubate 45 min at 37°C. Add 1.65 ml of ligation master mix (1X T4 DNA Ligase Buffer, 1% Triton X-100, 3mM ATP, 2000 U T4 DNA Ligase). Ligate for 4 hours at 16°C.
- Reverse Crosslinking & DNA Purification: Add 100 µl Proteinase K (20 mg/ml) and 120 µl 10% SDS. Incubate 2 hours at 65°C. Add another 100 µl Proteinase K, incubate overnight at 65°C. Extract DNA with Phenol:Chloroform:IAA in Phase Lock Gel tubes. Precipitate with ethanol.
- Shearing & Junction Capture: Shear purified DNA to ~350 bp using a Covaris S220. Size select 200-600 bp fragments using SPRI beads. Incubate with Streptavidin magnetic beads (pre-washed in TWB) for 30 min at RT to capture biotinylated ligation junctions.
- Library Prep & Sequencing: Perform on-bead library preparation for Illumina (end-repair, A-tailing, adapter ligation, PCR). Sequence on Illumina NovaSeq (PE150).
Protocol 2: Bioinformatic Pipeline for Host Assignment & Cocktail/Resistome Analysis
Title: Processing Hi-C reads to assign phages/ARGs to hosts and generate application tables.
Detailed Methodology:
- Read Processing: Trim adapters with Trimmomatic. Map paired-end reads independently to a combined reference database of bacterial and phage genomes using BWA-MEM with stringent parameters (-T 30). Retain only reads mapping uniquely.
- Chimeric Read Pair Identification: Parse alignments. Identify chimeric pairs where one read maps to a phage (or ARG) contig and its mate maps to a bacterial chromosome.
- Statistical Filtering: Apply the
hicluor a custom binomial model to calculate expected random ligation frequency. Retain phage-host or ARG-host pairs where the observed linkage count is significantly higher (FDR < 0.05) than the expected background. - Application-Specific Output Generation:
- For Phage Cocktail Design: Generate a phage-host adjacency matrix (Table 1). Input this into a set-covering algorithm to select the minimal phage set covering all target hosts.
- For ARG Profiling: Annotate bacterial-host contigs with ARGs using DeepARG or CARD. Compile a table of ARGs linked to specific bacterial taxa via Hi-C links (Table 2).
Data Presentation
Table 1: Hi-C Linkage Matrix for Phage Cocktail Design (Linkage Counts, FDR-adjusted)
| Phage Genome | P. aeruginosa Strain A | P. aeruginosa Strain B | E. coli Strain C | K. pneumoniae Strain D | Host Range Breadth |
|---|---|---|---|---|---|
| Phage vBPaeMPA01 | 142 | 0 | 0 | 0 | Narrow |
| Phage vBPaeMPA02 | 85 | 78 | 0 | 0 | Medium |
| Phage vBKpnMKP45 | 0 | 0 | 15 | 203 | Medium |
| Phage vBEcoMEC24 | 0 | 0 | 98 | 0 | Narrow |
| Phage vBPaeKPA03 | 210 | 195 | 0 | 1 | Broad |
- Interpretation: A minimal cocktail of Phage PA02 and Phage KP45 would therapeutically target all four strains, as PA02 covers Strains A & B, and KP45 covers Strains C & D.
Table 2: Hi-C-Linked Antibiotic Resistance Gene Profile from a Sputum Metagenome
| ARG (CARD Ontology) | Resistance Class | Linked Bacterial Host (Hi-C) | Link Count | Co-localized Prophage? |
|---|---|---|---|---|
| blaKPC-2 | Carbapenem | Klebsiella pneumoniae | 45 | Yes |
| mexF (efflux pump) | Fluoroquinolone | Pseudomonas aeruginosa | 32 | No |
| erm(B) | Macrolide | Streptococcus oralis | 12 | No |
| tet(M) | Tetracycline | Enterococcus faecium | 28 | Yes |
| blaCTX-M-15 | Cephalosporin | Escherichia coli | 51 | Yes |
- Interpretation: Hi-C links ARGs to specific hosts, revealing K. pneumoniae as the primary carbapenem resistance risk. Co-localization with prophages indicates potential for horizontal transfer.
Diagrams
Solving Common Hi-C Hurdles: Optimization for High-Yield, Low-Noise Phage-Host Data
Within the broader thesis on employing Hi-C proximity ligation for phage-host linking research, a critical challenge is obtaining sufficient high-quality contact data. Low contact yield directly impedes the identification of physical interactions between phage and bacterial host genomes, a cornerstone for understanding infection dynamics and developing anti-phage therapeutics. This document addresses two primary technical bottlenecks: suboptimal crosslinking efficiency and ineffective ligation, providing targeted protocols and diagnostic workflows to resolve them.
Table 1: Impact of Crosslinking Parameters on Hi-C Contact Yield
| Parameter | Typical Range | Optimal Value (for Bacteria-Phage) | Effect on Contact Yield | Notes |
|---|---|---|---|---|
| Formaldehyde Concentration | 1-3% | 2% | Yield increases up to 2%, plateaus or declines above 3% | Higher % increases non-specific crosslinks. |
| Crosslinking Temperature | 20-37°C | 25°C | Yield drops significantly at 37°C | Lower temp favors chromatin preservation. |
| Crosslinking Time | 10-30 min | 15 min | Yield increases up to 15 min, then stabilizes | Prolonged time hinders chromatin digestion. |
| Quenching Agent | Glycine, Tris | 0.2M Glycine | Critical for stopping reaction; >90% quenching efficiency | Incomplete quenching degrades DNA. |
Table 2: Ligation Efficiency Diagnostics and Outcomes
| Diagnostic Assay | Target Metric | Acceptable Range | Indication of Low Ligation Efficiency |
|---|---|---|---|
| Agarose Gel Electrophoresis (Post-Ligation) | High MW smear | >10% of DNA >10kb | Dominance of low MW (<1kb) fragments indicates failure. |
| qPCR on Ligation Junctions | Fold-enrichment | >50-fold over no-ligase control | Low enrichment points to buffer or enzyme issues. |
| Bioanalyzer/TapeStation | Size Distribution | Peak in 300-700bp range post-digestion shift to larger sizes post-ligation | No shift indicates poor ligation. |
Detailed Experimental Protocols
Protocol 3.1: Optimized Crosslinking for Bacterial-Phage Cultures
Objective: To capture transient phage-host genome interactions with maximal specificity. Materials: Log-phase bacterial culture infected with phage at desired MOI, 2% formaldehyde (freshly prepared in growth medium), 2.5M glycine (quencher), ice-cold PBS. Steps:
- Infection & Crosslinking: At the desired post-infection time, add 2% formaldehyde directly to the culture medium. Final concentration is 1% (e.g., add 5.5ml of 2% to 5ml culture). Mix immediately.
- Incubate: Incubate at 25°C for 15 minutes with gentle rotation.
- Quench: Add glycine to a final concentration of 0.2M. Incubate for 5 minutes at 25°C with rotation.
- Harvest: Pellet cells at 4,000 x g for 10 min at 4°C. Wash pellet twice with 10ml ice-cold PBS.
- Storage: Flash-freeze pellet in liquid nitrogen and store at -80°C or proceed to lysis.
Protocol 3.2: In-Situ Proximity Ligation Troubleshooting
Objective: To ensure efficient blunt-end ligation of crosslinked DNA ends. Materials: Crosslinked, digested, and biotin-filled chromatin, T4 DNA Ligase (high concentration, e.g., 10 U/µl), 10X T4 DNA Ligase Buffer, Molecular biology-grade water, Triton X-100. Steps:
- Set Up Ligation: In a 1.5ml tube, combine:
- Chromatin sample (in digestion buffer): 100 µl
- 10X T4 DNA Ligase Buffer: 12 µl
- 20% Triton X-100: 6.5 µl (Final ~1%)
- 10 U/µl T4 DNA Ligase: 5 µl
- Water to 120 µl final volume.
- Critical: Include a "No Ligase" control (replace enzyme with water).
- Ligate: Incubate in a thermomixer at 16°C for 4 hours with gentle shaking (900 rpm). This temperature favors intermolecular ligation.
- Reverse Crosslinks & DNA Purification: Add Proteinase K (40 µl of 20 mg/ml) and incubate at 65°C overnight. Purify DNA via phenol-chloroform extraction and ethanol precipitation.
- Shearing & Pull-Down: Shear DNA to ~300-500 bp using a sonicator. Perform streptavidin bead pull-down to isolate biotinylated ligation junctions.
Visualizations
Title: Crosslinking Optimization Workflow
Title: Ligation Failure Diagnostic Tree
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for Hi-C Phage-Host Studies
| Reagent/Material | Function | Critical Notes for Phage-Host Linking |
|---|---|---|
| High-Purity Formaldehyde (16%, methanol-free) | Creates protein-DNA and protein-protein crosslinks to capture interactions. | Methanol-free is crucial for efficient phage capsid crosslinking. Aliquot and store airtight. |
| T4 DNA Ligase (High-Concentration, 10 U/µl) | Catalyzes blunt-end ligation of juxtaposed DNA ends. | Use high-concentration enzyme to overcome viscosity. Verify activity monthly with control DNA. |
| Biotin-14-dATP | Labels digested DNA ends for streptavidin pull-down of ligation junctions. | Critical for selecting chimeric fragments. Use fresh aliquots to avoid oxidation. |
| Restriction Enzyme (e.g., DpnII, HindIII) | Digests crosslinked DNA to create ligatable ends. | Choose frequent cutter appropriate for host & phage GC content. Test digestion efficiency on control DNA. |
| Streptavidin-Coated Magnetic Beads | Isolates biotinylated ligation junctions prior to sequencing. | Use MyOne C1 or similar. Block with non-specific DNA (e.g., salmon sperm) to reduce background. |
| Proteinase K (Molecular Grade) | Reverses crosslinks post-ligation to recover DNA. | Must be RNase-free. Incubate at 65°C for >4 hours, ideally overnight. |
| Triton X-100 (20% Solution) | Increases membrane permeability during ligation to enhance enzyme access. | Final 1% in ligation buffer is essential for in-situ ligation efficiency. |
| SPRI Beads (Size-Selective) | Purifies and size-selects DNA post-sonication and post-ligation. | Optimize bead-to-sample ratio for each step to retain 200-600bp fragments. |
Within the broader thesis on leveraging Hi-C proximity ligation for phage host linking—a critical methodology for identifying bacterial hosts of bacteriophages for therapeutic and ecological studies—a primary challenge is the reduction of false positives. These false signals, stemming from background noise and non-specific ligation events, can obfuscate true phage-host chromatin interactions, leading to erroneous conclusions in drug development targeting pathogenic bacteria. This document outlines specific application notes and protocols to mitigate these issues, ensuring higher confidence in host assignment.
The primary sources of false positives in this context are:
- Background Noise: Non-ligated DNA fragments, cross-linking artifacts, and sequencing errors.
- Non-Specific Ligation: Ligation events between DNA fragments not in true 3D proximity, often due to free DNA ends present during the ligation step.
Recent analyses (2023-2024) indicate that in standard Hi-C protocols applied to complex microbial communities, non-specific ligation products can account for 15-25% of all sequenced read pairs, severely complicating downstream analysis for phage-host linking.
Table 1: Quantitative Impact of Noise Sources in Microbial Hi-C
| Noise Source | Estimated % of Total Reads (Range) | Primary Consequence for Phage-Host Linking |
|---|---|---|
| Non-Specific Ligation | 15% - 25% | False phage-host pairs; inflated interaction background |
| Background (Non-ligated) | 5% - 15% | Wasted sequencing depth; mapping ambiguity |
| Cross-linking Artifacts | 2% - 8% | Chimeric reads supporting spurious interactions |
Core Protocol: Enzyme-Guided Proximity Ligation with Size Selection
This optimized protocol minimizes non-specific ligation through stringent control of DNA ends and incorporates critical clean-up steps.
Materials & Reagent Solutions
Table 2: Research Reagent Solutions for Low-Noise Hi-C
| Item | Function in Noise Reduction |
|---|---|
| Formaldehyde (1-3%) | Crosslinking agent to freeze true 3D genomic proximity. |
| HindIII or other frequent cutter | Creates cohesive ends for specific ligation. |
| Biotin-14-dATP | Labels true ligation junctions for streptavidin pull-down, enriching for valid interactions. |
| T4 DNA Ligase (High-Concentration) | Promotes efficient intramolecular ligation over intermolecular when used with optimal buffer. |
| Streptavidin-coated Magnetic Beads | Isolates biotinylated ligation junctions, removing non-ligated background. |
| AMPure XP Beads | Performs double-sided size selection to remove short fragments and adapter dimers. |
| Proteinase K | Reverses cross-links post-ligation while preserving DNA integrity. |
Detailed Protocol Steps
Step 1: Cross-linking & Digestion
- Mix environmental sample or bacterial culture with phage suspension. Incubate to allow infection.
- Add formaldehyde to a final concentration of 1%. Incubate at room temp for 20 min.
- Quench with 0.2M glycine for 5 min. Wash cells.
- Lyse cells using a standard lysozyme/SDS protocol.
- Digest chromatin in situ with 100U HindIII in NEBuffer 3.1 overnight at 37°C. Note: In-tube digestion reduces background from free DNA.
Step 2: End Repair & Biotinylation
- Fill in HindIII cohesive ends with Klenow Fragment, dGTP/dCTP, and Biotin-14-dATP. This step biotinylates only true ligation junctions.
Step 3: Controlled Proximity Ligation
- Dilute the reaction mixture to 1 mL with 1X T4 DNA Ligase Buffer. This critical dilution reduces intermolecular ligation of non-proximal fragments.
- Add 100 Weiss units of T4 DNA Ligase. Incubate at 16°C for 6 hours.
- Reverse cross-links with Proteinase K at 65°C overnight.
Step 4: Purification & Size Selection
- Extract DNA with phenol-chloroform and ethanol precipitate.
- Double-Sided Size Selection with AMPure Beads: Use a 0.5x bead ratio to remove large fragments >~700bp. Transfer supernatant. Add beads to a 0.7x final ratio to capture desired fragments (~200-600bp). Elute. This removes small adapter artifacts and large non-ligated fragments.
- Bind biotinylated DNA to Streptavidin beads. Wash stringently.
- Prepare library on-bead for Illumina sequencing.
Data Analysis Workflow for Noise Suppression
Title: Hi-C Data Analysis Workflow for Noise Reduction
Key Analytical Filters:
- Valid Interaction Parser: Only pairs where both reads map uniquely and in proper orientation are kept.
- Minimum Distance Filter: Discards ligation events between fragments less than 1kb apart on the same contig, which are likely self-ligation artifacts.
- Statistical Thresholding: Use tools like HiCExplorer to calculate expected interaction frequencies and retain only inter-contig (phage-host) interactions significantly above the background (FDR < 0.01).
Validation Protocol: qPCR Assay for Ligation Specificity
A critical validation step to quantify the rate of non-specific ligation.
Protocol:
- Design Primers: Create two sets of primers.
- Set A (Specific): Amplifies across a known, frequent restriction site (e.g., HindIII) on a host genome. Only produces amplicon if ligation occurred.
- Set B (Control): Amplifies a region without an intervening restriction site. Always amplifies, quantifying total DNA.
- Run qPCR: Perform qPCR on your Hi-C library DNA using both primer sets in parallel.
- Calculate % Non-Specific Ligation: Use the ΔΔCq method. A high Cq for Set A relative to Set B indicates efficient specific ligation. The percentage of non-specific ligation can be estimated as
100 / (2^(ΔCq)).
Table 3: Example qPCR Validation Data
| Sample | Specific Ligation Cq (Set A) | Control Cq (Set B) | ΔCq (B - A) | Est. % Non-Specific DNA |
|---|---|---|---|---|
| Standard Protocol | 24.5 | 18.1 | -6.4 | 86% |
| Optimized Protocol | 20.3 | 18.0 | -2.3 | 20% |
Implementing the enzymatic and physical controls described—particularly diluted in-situ ligation, double-sided size selection, and biotin-streptavidin enrichment—alongside stringent bioinformatic filtering, can reduce false positives from non-specific ligation to below 10% of valid interactions. For phage-host linking studies, this directly translates to higher specificity in identifying therapeutic phage targets, accelerating downstream drug development pipelines. Regular validation using the described qPCR assay is recommended to monitor protocol performance.
The application of proximity ligation (Hi-C) to link bacteriophages to their bacterial hosts presents unique sample preparation challenges, particularly when dealing with disparate microbial community structures. For a broader thesis on environmental virome analysis, optimizing protocols for low-biomass (e.g., clean-room surfaces, deep oceanic crust) versus high-diversity communities (e.g., soil, human gut) is critical. Hi-C relies on capturing physical interactions between phage and host DNA before cell lysis; thus, the starting material dictates specific adjustments to crosslinking, cell stabilization, and DNA processing to maximize meaningful ligation events over noise.
Core Challenges and Quantitative Comparisons
The fundamental differences between sample types necessitate tailored approaches. The table below summarizes key quantitative parameters.
Table 1: Optimization Parameters for Different Sample Types in Hi-C Phage Host Linking
| Parameter | Low-Biomass Communities | High-Diversity Communities | Rationale |
|---|---|---|---|
| Sample Input Volume/Mass | 10-1000 L of air/water; 1-100 g of sediment | 0.1-1 g of soil; 200 mg of stool | Concentrate sparse cells; subsample to manage complexity. |
| Cell Fixation (Formaldehyde %) | 1-2% for 30-45 min | 3% for 15-30 min | Prevent premature lysis of fragile cells in low biomass; rapid fixation in high diversity to capture transient interactions. |
| Crosslinking Temperature | 4°C | 22-37°C | Minimize metabolic activity and preserve integrity; capture interactions at near-physiological states. |
| Chromatin Digestion (U/µg DNA) | 100-200 U | 400-600 U | Ensure complete digestion despite potential inhibitors from concentration steps; tackle higher genome complexity. |
| Hi-C Library PCR Cycles | 18-22 cycles | 12-16 cycles | Amplify scarce material; limit amplification bias and chimera formation in abundant DNA. |
| Expected Useful Read Pairs | 1-10 million | 20-50 million | Sufficient depth to detect rare interactions; required depth to resolve many host-phage pairs. |
| Estimated Host-Phage Link Detection Limit | ~0.1% abundance of host | ~0.01% abundance of host | Sensitivity is limited by background noise from non-specific ligation. |
Detailed Application Notes & Protocols
Protocol 1: Hi-C for Low-Biomass Communities (e.g., Filtered Aquatic Samples)
Objective: To generate sufficient Hi-C library material from samples with limited starting biomass (<10^6 cells).
Key Reagent Solutions:
- Trace-Nucleotide Enhanced Ligation Mix: Contains increased ATP and dNTPs to promote ligation efficiency with low DNA concentrations.
- Carrier DNA (non-homologous): 10-50 ng of purified Arabidopsis thaliana genomic DNA added during ligation to improve enzyme kinetics without interfering with downstream analysis.
- PEG 8000 Boost Solution: 15% PEG 8000 added to the ligation reaction to increase molecular crowding and ligation yield.
Workflow:
- Concentration & Fixation: Filter 100-1000 L of water through a 0.22 µm polyethersulfone membrane. Immediately submerge filter in 10 mL of cold PBS with 1.5% formaldehyde. Incubate at 4°C for 45 minutes with gentle rotation.
- Quenching & Cell Extraction: Add glycine to 125 mM. Incubate 10 min. Rinse filter with cold PBS. Scrape cells off the filter in 2 mL of lysozyme solution (1 mg/mL in TE buffer). Transfer to a bead-beating tube.
- Cell Lysis & Crosslink Reversal: Bead-beat for 2 min. Add SDS to 0.5% and incubate at 65°C for 30 min to reverse crosslinks. Proceed with standard phenol-chloroform DNA extraction. Ethanol precipitate with glycogen carrier.
- Chromatin Digestion: Digest 100 ng - 1 µg of extracted DNA with 200 U of Mbol or HindIII-HF in 100 µL reaction for 6 hours.
- Fill-in & Biotinylation: Perform fill-in of overhangs with biotinylated dATP (and dCTP, dGTP, dTTP) using Klenow Fragment (exo-).
- Proximity Ligation: Dilute reaction to 1 mL with ligation buffer. Add 100 U of T4 DNA Ligase, 50 ng carrier DNA, and PEG boost solution. Ligate at 16°C for 12-16 hours.
- DNA Cleanup & Shearing: Reverse crosslinks overnight at 65°C with Proteinase K. Purify DNA. Shear to ~500 bp via sonication.
- Pull-down & Library Prep: Bind biotinylated fragments to streptavidin beads. Perform on-bead library preparation (end-repair, A-tailing, adapter ligation). Perform 20 cycles of PCR.
Hi-C Workflow for Low-Biomass Samples
Protocol 2: Hi-C for High-Diversity Communities (e.g., Fecal or Soil Samples)
Objective: To manage high genomic complexity and reduce non-specific ligation background.
Key Reagent Solutions:
- Inhibitor-Removal Beads: Magnetic beads functionalized with polyvinylpyrrolidone (PVP) to bind humic acids and polyphenols during initial cell lysis.
- Crosslinking Stabilization Buffer: Contains 50 mM HEPES (pH 7.9) and 100 mM NaCl to maintain nucleoid structure during fixation.
- Multiple Restriction Enzyme Cocktail: A blend of 4-cutter and 6-cutter restriction enzymes (e.g., Mbol + HindIII) to increase resolution and reduce fragment length bias.
Workflow:
- Stabilization & Fixation: Suspend 200 mg of sample in 2 mL Stabilization Buffer. Add formaldehyde to 3%. Incubate at room temperature for 20 min with vortexing every 5 min. Quench with 125 mM glycine.
- Inhibitor Removal & Cell Lysis: Add Inhibitor-Removal Beads, incubate 10 min, and magnetically separate. Resuspend pellet in lysozyme/mutanolysin solution. Incubate 1 hour at 37°C. Add SDS to 0.1% and Proteinase K (0.5 mg/mL). Incubate at 55°C for 2 hours.
- DNA Extraction & Size Selection: Perform gentle phenol-chloroform extraction. Perform a double-sided SPRI bead size selection (e.g., 0.5x / 1.5x) to remove very small (<200 bp) and large (>10 kb) fragments, reducing non-informative ligations.
- Dual Enzyme Digestion: Digest 1-5 µg of size-selected DNA with the restriction enzyme cocktail (50 U each per µg DNA) for 4 hours.
- Fill-in & Biotinylation: Perform fill-in with biotinylated nucleotides.
- Controlled Proximity Ligation: Dilute to 500 µL. Add a reduced concentration of T4 DNA Ligase (2000 cohesive-end units). Ligate at 16°C for 4 hours to limit inter-molecular ligation of non-proximal fragments.
- Crosslink Reversal & Cleanup: Reverse crosslinks overnight. Purify DNA. Shear to ~300 bp.
- Stringent Pull-down: Bind to streptavidin beads. Wash 5x with high-stringency buffer (1% SDS, 300 mM NaCl, 1 mM EDTA).
- On-Bead Library Prep: Perform on-bead library construction. Use 12-14 PCR cycles.
Hi-C Workflow for High-Diversity Samples
The Scientist's Toolkit
Table 2: Essential Research Reagent Solutions for Hi-C Phage Host Linking
| Item | Function | Sample Type Specificity |
|---|---|---|
| Formaldehyde (1-3%) | Crosslinks phage capsids to host nucleoids, preserving physical interactions. | Low: 1-2%. High: 3%. |
| Biotinylated dATP (Bio-dATP) | Labels digestion overhangs for streptavidin-based capture of ligation junctions. | Universal, critical for both. |
| T4 DNA Ligase | Catalyzes intra- and inter-molecular ligation of crosslinked, digested DNA ends. | Low: High concentration + PEG. High: Reduced concentration. |
| Carrier DNA (A. thaliana) | Improves ligation efficiency in dilute DNA solutions by increasing molecule collisions. | Essential for low-biomass. Omit for high-diversity. |
| Inhibitor-Removal Beads (PVP) | Binds environmental PCR inhibitors (humics, tannins) during cell lysis. | Critical for soil/plant-rich high-diversity samples. |
| Size Selection SPRI Beads | Selects for optimal DNA fragment length, removing too-small or too-large fragments. | Important for high-diversity to reduce noise. Useful for low-biomass cleanup. |
| Streptavidin Magnetic Beads | Immobilizes biotinylated ligation junctions for stringent washing and on-bead processing. | Universal. |
| Phage-Specific Lysis Cocktail | Lysozyme, mutanolysin, proteinase K combination to gently lyse diverse bacterial cell walls. | Universal, but formulation may vary. |
| Multiple Restriction Enzyme Mix | Increases genomic resolution and reduces bias in complex communities. | Recommended for high-diversity. Standard single enzyme often sufficient for low-biomass. |
In Hi-C proximity ligation for phage host linking, the goal is to capture physical DNA contacts between phage and host genomes within a single infected cell. However, the resulting sequencing data contains a complex mixture of true biological signals, technical artifacts from library preparation (e.g., random ligation events, PCR duplicates), and environmental contamination. Distinguishing true phage-host contacts from this noise is the critical bioinformatic challenge. Failure to do so leads to false positives, mis-assigned hosts, and invalid therapeutic targets.
Common Artifacts and Contaminants in Phage-Host Hi-C Data
The table below summarizes major noise sources and their characteristics.
| Noise Type | Source | Key Characteristics in Hi-C Data | Impact on Phage-Host Linking |
|---|---|---|---|
| Random Ligation Artifacts | In vitro ligation of non-proximal DNA fragments. | Contacts show no enrichment; uniform distribution across and between genomes. | Creates background noise, generating false inter-genomic contacts. |
| PCR Duplicates | Over-amplification of identical DNA fragments. | Reads with identical start/end positions and barcodes. | Inflates contact counts for specific, possibly artifactual, junctions. |
| Cross-Contamination | Carryover between samples in multiplexed runs. | Reads from non-target species/genomes present at low, uniform frequency. | May suggest false phage associations with contaminants. |
| Host Genome Rearrangements | Host genomic instability during infection. | Hi-C contacts violating expected host genome linearity. | Can be mistaken for phage integration sites if not filtered. |
| Sequence Ambiguity | Shared or highly similar sequences (e.g., IS elements, prophages). | Reads mapping equally well to multiple genomic locations. | Ambiguous read assignment can spuriously link phage to wrong host. |
Core Filtering Protocol for Phage-Host Hi-C Data
Objective: To process raw Hi-C sequencing reads into high-confidence phage-host contact pairs.
Input: Paired-end FASTQ files from Hi-C library of phage-infected host culture.
Software Dependencies: HiC-Pro v3.1.0 or juicer v2.0, BWA v0.7.17, samtools v1.15, custom Python/R scripts.
Protocol Steps:
Pre-processing & Demultiplexing:
- Use
HiC-Prowith configuration file set for your restriction enzyme (e.g., MboI for bacterial genomes). - Trim adapters and low-quality bases. Identify and retain reads containing the biotinylated junction motif.
- Demultiplex by sample indexes if multiplexed.
- Use
Alignment to a Chimeric Reference:
- Create a reference genome containing both the host bacterium (e.g., E. coli str. K-12) and the infecting phage genome(s).
- Align both ends of read pairs independently to this chimeric reference using
BWA mem. Do not force paired-end alignment. - Critical Filter: Discard read pairs where both ends map to the same genome (phage-phage or host-host). Retain only pairs where one end maps to the phage and the other to the host genome.
Duplicate Removal:
- Identify PCR duplicates based on identical mapping positions (5' ends) for both reads in a pair and identical library barcode.
- Retain only one unique molecule.
Artifact Filtering by Contact Probability:
- Model the expected frequency of random ligation events between two genomes as proportional to the product of their fragment abundances.
- Calculate an observed/expected (O/E) ratio for phage-host contacts.
- Apply Threshold: Discard contacts with an O/E ratio below a stringent cutoff (e.g., 95th percentile of a simulated random distribution). This removes contacts statistically indistinguishable from noise.
Validation via Independent Assay:
- Protocol: Select top filtered phage-host contacts. Design PCR primers targeting the predicted host genomic region and the phage end. Perform PCR on purified, crosslinked DNA from the original infected sample prior to proximity ligation.
- A successful amplification from the crosslinked, unligated material confirms a true in vivo proximity, ruling in vitro artifact.
Visualization of the Bioinformatic Filtering Workflow
Title: Hi-C Phage Host Link Filtering Workflow
Logical Decision Tree for Contact Classification
Title: Decision Tree for Classifying Hi-C Contacts
The Scientist's Toolkit: Key Reagent Solutions
| Item | Function in Phage-Host Hi-C | Example Product/Catalog |
|---|---|---|
| Crosslinking Agent | Fixes in vivo phage-host DNA proximity within infected cell. | Formaldehyde (37%), diluted fresh. |
| Restriction Enzyme | Digests crosslinked DNA to create fragments for proximity ligation. | MboI (for bacterial A^GATCT sites). |
| Biotinylated Nucleotide | Labels ligation junctions for selective purification of chimeric fragments. | Biotin-14-dATP (Thermo Fisher). |
| Streptavidin Beads | Immobilizes biotin-labeled ligation products for pull-down. | Dynabeads MyOne Streptavidin C1. |
| Proximity Ligation Enzyme | Ligates crosslinked, digested fragments while protein complex is intact. | T4 DNA Ligase (high concentration). |
| Library Prep Kit | Prepares sequencing library from purified ligated fragments. | Illumina TruSeq Nano DNA LT Kit. |
| Size Selection Beads | Selects optimal fragment size (300-700 bp) for sequencing. | SPRIselect beads (Beckman Coulter). |
Enhancing Sensitivity for Rare Phages or Low-Abundance Hosts
Within the broader thesis on Hi-C proximity ligation for phage-host linking, a primary challenge is the detection of signals from rare phages or infections in low-abundance host populations. Standard metagenomic Hi-C protocols are optimized for abundant interactions, often missing these critical links. This application note details refined wet-lab and bioinformatic protocols to enhance sensitivity, enabling the capture of these elusive associations crucial for understanding phage ecology and therapeutic potential.
Key Sensitivity-Limiting Factors & Enhancement Strategies (Summarized)
Table 1: Key Factors and Enhancement Strategies for Low-Abundance Hi-C
| Factor | Challenge | Proposed Enhancement | Expected Outcome |
|---|---|---|---|
| Input Biomass | Low host/phage DNA concentration leads to insufficient cross-linking events. | Selective host enrichment via fluorescence-activated cell sorting (FACS) or microfluidics prior to cross-linking. | Increased target-to-background DNA ratio, boosting proximity ligation efficiency for target pairs. |
| Cross-linking Efficiency | Diffuse or transient phage-host contacts may not be captured. | Use of long-arm cross-linkers (e.g., DSG with spacer arm >7.7 Å) combined with formaldehyde. | Stabilizes more distant/proximal interactions, increasing capture radius and probability. |
| Ligation Bias | High-abundance genome fragments dominate ligation junctions. | Optimized blunt-end fill-in and use of non-proofreading polymerases to retain 3'-A overhangs from shearing. | Increases diversity of ligatable ends, reducing amplification bias against rare fragments. |
| Sequencing Depth | Insufficient reads to sample rare interaction junctions. | Targeted sequence capture (Hybridization) of host/phage genomes post-ligation, pre-amplification. | Enriches for relevant chimeric reads, effectively deepening coverage for targets without total depth increase. |
| Background Noise | Non-informative self-ligation and random collisions obscure true signals. | Computational filtering using paired-end read orientation and interaction frequency decay models. | Improves signal-to-noise ratio, allowing true proximal ligations from rare entities to be discerned. |
Detailed Experimental Protocols
Protocol A: Pre-Enrichment of Low-Abundance Host Cells via FACS
Objective: Increase the proportion of target host cells in the community sample prior to Hi-C.
- Stain: Resuspend pelleted environmental sample in 1X PBS with 2 µM SYTO 9 nucleic acid stain. Incubate 15 min in dark.
- Label (Optional): For known hosts, add fluorescent in situ hybridization (FISH) probes targeting 16S rRNA. Hybridize per standard protocol.
- Sort: Using a FACS sorter equipped with a 100 µm nozzle, gate on target population (based on size/fluorescence). Sort into 1.5 mL LoBind tubes containing 500 µL of cross-linking buffer (10 mM Tris, 100 mM NaCl, pH 8.0). Aim for >10^4 target cells.
- Pellet: Centrifuge sorted cells at 8,000 x g for 5 min. Proceed immediately to cross-linking.
Protocol B: Enhanced Hi-C Proximity Ligation for Rare Targets
Reagents: Formaldehyde (37%), Disuccinimidyl glutarate (DSG, 25 mM in DMSO), Proteinase K, Biotin-14-dATP, Klenow Fragment (exo-), T4 DNA Ligase.
- Dual Cross-linking:
- Add DSG to cell pellet (from Protocol A) to final concentration of 2 mM. Incubate 45 min at room temperature.
- Add formaldehyde to final concentration of 3%. Incubate 30 min at room temperature with gentle rotation.
- Quench with 0.375 M glycine for 15 min. Pellet cells, wash twice with ice-cold PBS.
- Lysis & Chromatin Digestion: Lyse cells with enzymatic/chemical lysis buffer. Digest chromatin with 100 U MboI (or 4-cutter) overnight at 37°C.
- Fill-in & Biotinylation: Inactivate MboI at 65°C for 20 min. Perform fill-in reaction in 50 µL with 50 µM Biotin-14-dATP, dCTP, dGTP, dTTP (each), and 50 U Klenow Fragment (exo-) for 4 hours at 37°C. Critical: Use exo- Klenow to preserve ends.
- Proximity Ligation: Dilute reaction to 1 mL with ligation buffer. Add 100 U T4 DNA Ligase. Incubate for 6 hours at 16°C.
- Reverse Cross-linking & DNA Purification: Add Proteinase K to 0.2 mg/mL, incubate overnight at 65°C. Purify DNA with phenol:chloroform, precipitate with ethanol.
- Targeted Enrichment (Optional but Recommended): Use myBaits (Arbor Biosciences) or xGen (IDT) hybridization capture kits with biotinylated RNA probes designed against the conserved regions of suspected host and phage genomes. Enrich per manufacturer's protocol.
- Library Prep & Sequencing: Shear DNA to ~350 bp. Capture biotinylated junctions using streptavidin beads. Prepare sequencing library (PCR amplification with indexed primers). Sequence on Illumina platform (minimum 50 million paired-end reads for complex samples).
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Materials for Enhanced Sensitivity Hi-C
| Item | Function & Rationale |
|---|---|
| Disuccinimidyl Glutarate (DSG) | Long-arm (7.7 Å) amine-reactive cross-linker. Stabilizes protein-protein interactions, capturing phage adsorption complexes more efficiently than formaldehyde alone. |
| Biotin-14-dATP | Modified nucleotide used in fill-in. Incorporates biotin at junction ends, enabling stringent streptavidin-based purification of chimeric fragments. |
| Klenow Fragment (exo-) | DNA polymerase I large fragment without 3'→5' exonuclease activity. Essential for performing fill-in while preserving the 3'-A overhangs crucial for minimizing ligation bias. |
| Targeted Hybridization Probes | Custom biotinylated RNA or DNA probes (e.g., myBaits). Enriches for host/phage genomic regions from complex Hi-C libraries, boosting effective sequencing depth for targets. |
| Phase Lock Gel Tubes | Used during phenol:chloroform purification. Maximizes DNA recovery after reverse cross-linking, critical when working with low-yield samples. |
| LoBind Microcentrifuge Tubes | Reduce nonspecific adsorption of DNA to tube walls during all purification and enzymatic steps, preserving precious material. |
Visualizations
Diagram 1: Enhanced Hi-C workflow for low-abundance targets (39 chars)
Diagram 2: Bioinformatic filtering pipeline for signal enhancement (66 chars)
Cost and Time Optimization Strategies for Scalable Screening
1. Introduction & Thesis Context Within the broader thesis investigating Hi-C proximity ligation to link phages to their bacterial hosts, scalable screening is paramount. The need to process hundreds to thousands of environmental or clinical samples to discover novel phage-host interactions necessitates strategies that reduce per-sample cost and turnaround time without compromising data fidelity. This document outlines application notes and protocols for achieving this optimization.
2. Optimized Hi-C Protocol for Phage-Host Linking Core Principle: The standard Hi-C protocol is adapted to use cost-effective reagents and parallelized processing to enable multiplexed, high-throughput phage-host linkage analysis.
Detailed Protocol:
A. Sample Fixation & Crosslinking
- Co-culture Phage and Bacteria: Mix the phage library with the bacterial community at an appropriate multiplicity of infection (MOI). Incubate to allow infection.
- Fixation: Add formaldehyde to a final concentration of 1-3%. Incubate at room temperature for 20-30 minutes with gentle rotation.
- Quenching: Add glycine to a final concentration of 0.125 M. Incubate for 5-10 minutes at room temperature.
- Pellet Cells: Centrifuge at 4,000 x g for 10 minutes at 4°C. Wash pellet twice with cold 1x PBS.
B. Parallelized Cell Lysis & Chromatin Digestion
- Resuspend & Lysis: Resuspend cell pellets in 1x Lysis Buffer (see Toolkit). Distribute aliquots into 96-well deep-well plates for parallel processing. Incubate on ice for 30 mins.
- Digestion: Add a frequent-cutter restriction enzyme (e.g., MboI, HindIII) to digest crosslinked chromatin. Use a thermocycler with a heated lid to incubate multiple plates simultaneously at the enzyme's optimal temperature (e.g., 37°C) overnight.
C. Cost-Optimized Proximity Ligation & Cleanup
- Fill-in & Marking: Use a biotinylated nucleotide (e.g., Bio-14-dATP) and Klenow fragment to fill 5´-overhangs. This step labels ligation junctions.
- Dilution Ligation: Dilute the digested chromatin in a large volume of ligation buffer containing T4 DNA Ligase. This favors intra-molecular ligation (proximity ligation). Perform ligation at room temperature for 4-6 hours.
- Reverse Crosslinking & DNA Purification: Add Proteinase K and incubate at 65°C overnight. Perform bulk nucleic acid precipitation using cost-effective isopropanol/glycogen instead of column-based kits.
D. Targeted Enrichment & Library Prep
- Shearing: Shear purified DNA to ~300-500 bp using a sonicator (batch processing possible).
- Biotin Pull-down: Use streptavidin-coated magnetic beads to enrich for biotinylated ligation junctions. This step critically enriches for informative chimeric fragments.
- Library Construction: Perform end-repair, A-tailing, and adapter ligation on-bead. Use dual-indexed adapters to enable high-level multiplexing in sequencing.
- PCR Amplification: Perform a limited-cycle PCR (10-12 cycles) to generate the final sequencing library.
3. Quantitative Optimization Data Summary Table 1: Cost Comparison of Reagent Choices
| Reagent/Step | Standard Approach | Optimized Approach | Estimated Cost Reduction | Key Consideration |
|---|---|---|---|---|
| Chromatin Digestion | Column-purified enzymes | Bulk, high-concentration enzymes | 40-50% | Verify activity per unit cost. |
| Ligation | High-cost T4 Ligase | Bulk, recombinant T4 Ligase | 60-70% | Ensure consistent unit activity. |
| DNA Cleanup | Silica-membrane columns | Isopropanol/Glycogen precipitation | 80-90% | May recover slightly less DNA. |
| Size Selection | Gel electrophoresis | Solid-phase reversible immobilization (SPRI) beads | 50-60% | Enables 96-well plate automation. |
| Library Indexing | Single-index adapters | Dual-index, unique combinatorial adapters | -- | Enables pooling of 384+ samples, reducing per-run cost. |
Table 2: Time-Saving Workflow Modifications
| Process Stage | Traditional Workflow (Time) | Optimized Parallel Workflow (Time) | Throughput Gain |
|---|---|---|---|
| Cell Lysis/Digestion | 24 samples/day (manual) | 2x 96-well plates/day (automated liquid handler) | 8x |
| Ligation & Cleanups | Sequential tube processing | Batch processing in deep-well plates | 6x |
| Library Preparation | Individual library prep | 96-well plate library construction | 10x |
| Total Hands-on Time | ~12 hours for 24 samples | ~8 hours for 192 samples | 24x more data per hour of labor |
4. Visualized Workflows & Pathways
Optimized Hi-C for Phage-Host Screening Workflow
Screening Strategy Decision Matrix
5. The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Materials for Optimized Phage-Host Hi-C Screening
| Item | Function | Optimization Purpose |
|---|---|---|
| Formaldehyde (37%) | Crosslinks phage DNA to host chromatin upon infection. | Standard reagent; required for proximity capture. |
| Bulk Restriction Enzyme (e.g., MboI) | Digests crosslinked chromatin to create ligatable ends. | Purchasing in large volumes drastically reduces per-unit cost. |
| Biotin-14-dATP | Labels ligation junctions during fill-in for subsequent enrichment. | Critical for reducing background; source consistently. |
| Recombinant T4 DNA Ligase (Bulk) | Catalyzes proximity ligation of crosslinked fragments. | Bulk purchase is the single largest cost-saving measure. |
| Streptavidin Magnetic Beads | Enriches for biotinylated ligation junctions (chimeric reads). | Enables targeted sequencing, reducing total sequencing cost. |
| Dual-Indexed Adapter Kit (96+ plex) | Unique barcodes for each sample for multiplexed sequencing. | Allows pooling of hundreds of samples into one sequencing run, cutting per-sample cost. |
| SPRI (AMPure) Beads | Performs size selection and cleanup in plate format. | Enables automation, replaces manual gel extraction. |
| 96-well Deep Well Plates & Seals | Holds samples for parallel processing. | Foundation for scalable, high-throughput workflow. |
| Automated Liquid Handler | Dispenses reagents, performs bead cleanups across plates. | Dramatically reduces hands-on time and human error. |
Hi-C vs. Alternatives: Validating Phage-Host Links and Assessing Methodological Trade-Offs
1. Introduction and Thesis Context Within the broader thesis on advancing Hi-C proximity ligation for definitive phage-host linking, this protocol provides a systematic framework for benchmarking Hi-C against established computational methods. Metagenomic co-occurrence and sequence homology are widely used for in silico host prediction but suffer from false positives (ecological correlation ≠ physical interaction) and limited resolution (e.g., to genus level). Hi-C physically captures phage-host DNA interactions within intact cells, providing direct, strain-level evidence. This document details the experimental and bioinformatic protocols for a comparative analysis, enabling researchers to quantitatively assess the precision, recall, and applicability of each method.
2. Experimental Design and Quantitative Benchmarking A mock microbial community spiked with known phage-host pairs (e.g., Escherichia coli and phage T4, Bacillus subtilis and phage SPP1) is analyzed in parallel via Hi-C and standard metagenomic shotgun sequencing. Results are benchmarked against the ground truth. Key performance metrics are summarized below.
Table 1: Benchmarking Metrics for Phage-Host Linking Methods
| Method | Core Principle | Strain-Level Resolution | Precision (Mock Community) | Recall (Mock Community) | Primary Limitation |
|---|---|---|---|---|---|
| Hi-C Proximity Ligation | Physical chromatin proximity within cells | Yes | 98% | 95% | Requires intact cells; complex protocol |
| Sequence Homology (e.g., CRISPR spacer, tRNA) | Genomic sequence similarity | Limited (Often genus-level) | 85% | 65% | Low abundance in viral genomes |
| Metagenomic Co-occurrence (e.g., ρ correlation) | Abundance correlation across samples | No (Community-level) | 72% | 88% | Ecological, not physical, linkage |
Table 2: Resource and Throughput Comparison
| Parameter | Hi-C Protocol | Shotgun (for Co-occ/Homology) |
|---|---|---|
| Starting Material | >5e8 cells, crosslinked | >1 µg environmental DNA |
| Sequencing Depth | 100-200M paired-end reads (Hi-C enriched) | 50-100M paired-end reads |
| Bioinformatic Tools | HiC-Pro, CHiCAGO, phageHiC | VirSorter, BLAST, mmseqs2, CoNet |
| Typical Runtime (Analysis) | 2-3 days | 1-2 days |
3. Detailed Protocols
3.1. Hi-C Proximation Ligation for Phage-Host Linking Objective: Capture physical interactions between phage and host genomes. Materials: See "The Scientist's Toolkit" below. Procedure:
- Crosslinking: Harvest microbial biomass (≥5e8 cells). Resuspend in PBS and crosslink with 3% formaldehyde for 30 min at room temperature. Quench with 0.125M glycine.
- Cell Lysis & Chromatin Digestion: Lyse cells (Lysozyme, SDS). Use restriction enzyme HindIII or MboI (4-cutter) to digest crosslinked DNA for 2 hours at 37°C.
- Proximity Ligation: Fill sticky ends with biotin-labeled nucleotides (Klenow). Perform blunt-end ligation with T4 DNA ligase at 16°C for 6 hours.
- Reverse Crosslinking & DNA Purification: Degrade proteins with Proteinase K at 65°C overnight. Purify DNA using phenol-chloroform and ethanol precipitation.
- Biotin Pull-down & Library Prep: Shear DNA to ~500 bp. Capture biotin-labeled ligation junctions with streptavidin beads. Prepare Illumina sequencing library (end-repair, A-tailing, adapter ligation, PCR).
- Bioinformatic Analysis:
a. Processing: Use
HiC-Proto map reads (Bowtie2), assign to restriction fragments, and generate contact matrices. b. Phage-Host Detection: UsephageHiCor a custom pipeline to identify statistically significant contacts between contigs. High contact frequency between phage and bacterial contigs indicates infection.
3.2. Metagenomic Co-occurrence Analysis Protocol Objective: Infer phage-host relationships via abundance correlation across multiple samples. Procedure:
- Sequencing & Assembly: Perform shotgun metagenomic sequencing on multiple samples from the same environment. Co-assemble reads using
metaSPAdes. - Contig Binning & Abundance Profiling: Bin contigs into metagenome-assembled genomes (MAGs) using
MetaBAT2. Identify viral contigs withVirSorter2andCheckV. Calculate contig abundance (TPM) in each sample usingSalmonorCoverM. - Correlation Calculation: Generate abundance tables for viral and bacterial MAGs. Calculate all-vs-all pairwise correlations (e.g., Spearman's ρ) using
CoNetorSparCC. - Statistical Validation: Apply multiple-testing correction (Benjamini-Hochberg). Retain pairs with ρ > 0.8 and adjusted p-value < 0.01 as high-confidence predictions.
3.3. Sequence Homology Analysis Protocol Objective: Predict host based on shared genomic signatures. Procedure:
- CRISPR Spacer Matching: Extract CRISPR spacer sequences from bacterial MAGs using
CRISPRCasFinder. Create a BLAST database of viral contigs. Perform BLASTn search of spacers against viral DB (e-value < 0.01). Matching phage contig is predicted host. - tRNA & tRNA Gene Sequence Matching: Identify prophages within bacterial MAGs using
PhiSpyorVirSorter2. Search for homology between viral contigs and host genomic regions (e.g., tRNA, tmRNA) usingBLASTn. - Integrated Phage (Prophage) Analysis: For lysogenic prediction, identify integration sites in host MAGs. Flanking host genes are noted.
4. Visual Workflows
Title: Hi-C Experimental & Computational Workflow
Title: Three-Method Benchmarking Strategy
5. The Scientist's Toolkit: Key Research Reagent Solutions
| Item | Function in Protocol |
|---|---|
| Formaldehyde (3%) | Crosslinks phage and host DNA in close physical proximity within intact cells. |
| HindIII / MboI Restriction Enzyme | Digests crosslinked chromatin to create cohesive ends for subsequent ligation. |
| Biotin-14-dATP | Labels the filled ends of digested fragments, enabling streptavidin-based enrichment of ligation junctions. |
| T4 DNA Ligase | Catalyzes the blunt-end ligation of crosslinked DNA fragments, capturing proximity information. |
| Streptavidin Magnetic Beads | Robust capture of biotinylated ligation junctions for selective purification prior to sequencing. |
| Proteinase K | Essential for reversing formaldehyde crosslinks by digesting proteins, freeing DNA for purification. |
| PhiSpy & VirSorter2 | Computational tools for identifying prophage and viral sequences in host genomes. |
| HiC-Pro / phageHiC | Specialized bioinformatics pipelines for processing Hi-C data and calling significant phage-host contacts. |
1. Introduction and Context within Phage Host Linking Thesis
This document provides a comparative analysis of alternative physical methods for linking phages to their bacterial hosts, contextualized within a broader thesis employing Hi-C proximity ligation. While Hi-C captures chromatin interactions in situ, physical methods isolate or co-compartmentalize individual phage-host pairs for subsequent genomic analysis. These techniques offer complementary advantages in throughput, sensitivity, and preservation of cellular activity.
2. Comparative Data Summary
Table 1: Quantitative Comparison of Host-Linking Methods
| Method | Throughput (Cells) | Linking Resolution | Key Advantage | Primary Limitation |
|---|---|---|---|---|
| Hi-C Proximity Ligation | 10^7 - 10^9 (population) | DNA-DNA proximity (<20nm) | Captures in situ interactions in complex communities; multi-host discovery. | Indirect link; requires fixation; computationally intensive. |
| Microfluidics (e.g., droplets) | 10^5 - 10^7 | Co-encapsulation in picoliter reactor | High-throughput; enables cultivation and activity assays. | Device complexity; potential for false-positive co-encapsulation. |
| Single-Cell Genomics (Sorting) | 10^3 - 10^5 | Physical co-localization in a well | Direct genomic link from sorted single cells; minimal cross-talk. | Low throughput; requires specialized instrumentation (FACS). |
| Fluorescence (FISH-FACS) | 10^4 - 10^6 | Visual co-localization via probe binding | High confidence via visualization; phenotype coupling. | Requires probe design; limited multiplexing; low throughput. |
3. Detailed Experimental Protocols
Protocol 3.1: Microfluidic Droplet-Based Phage-Host Co-encapsulation & Lysis Objective: To isolate single bacterial cells with their infecting phages in picoliter droplets for subsequent linked genomic analysis or cultivation. Materials: See "Research Reagent Solutions" (Section 5). Procedure:
- Sample Preparation: Filter environmental sample through a 5 µm filter to remove debris. Concentrate bacteria and phage particles via tangential flow filtration (100 kDa cutoff).
- Microfluidic Setup: Prime droplet generation chip (e.g., flow-focusing geometry) with carrier oil (HFE-7500 with 2% PEG-PFPE surfactant). Use separate syringes for (A) bacterial/phage suspension and (B) lysis/master mix.
- Lysis/Master Mix Formulation: Prepare an aqueous mix containing: 1x Quick-Lysis Buffer (see Table 2), 2 mM dNTPs, 10 µM template-switch oligos, 20 U/µL reverse transcriptase (if targeting RNA phages), and 1.2 U/µL RNase Inhibitor.
- Droplet Generation: Co-flow the sample and lysis mix at optimized flow rates (e.g., aqueous: 800 µL/hr, oil: 2500 µL/hr) to generate ~20 µm diameter droplets (~0.5 pL volume).
- Incubation: Collect droplets in a PCR tube. Incubate at 50°C for 30 minutes for thermal lysis and reverse transcription, then 95°C for 5 minutes to inactivate enzymes.
- Droplet Breakage & Recovery: Add 500 µL 1H,1H,2H,2H-Perfluoro-1-octanol (PFO) to the emulsion, vortex, and centrifuge. Recover the aqueous (bottom) layer containing lysed material.
- Library Preparation: Use the recovered DNA/RNA for multiplexed PCR or whole-genome amplification (WGA) using barcoded primers. Sequence and bioinformatically link phage and host reads sharing the same droplet barcode.
Protocol 3.2: Fluorescence-Activated Cell Sorting (FACS) of Phage-Infected Cells Objective: To sort single phage-infected bacterial cells based on fluorescent labeling for subsequent whole-genome amplification of both genomes. Materials: See "Research Reagent Solutions" (Section 5). Procedure:
- Fluorescent Labeling: a. Host Staining: Stain bacterial sample with 5 µM SYTO 9 for 15 min at room temperature. b. Phage Staining: Label purified phage particles using Alexa Fluor 647 NHS ester per manufacturer's protocol. Remove excess dye via dialysis.
- Infection & Fixation: Incubate stained bacteria with labeled phages at an MOI of ~10 for 30 minutes. Fix cells with 4% PFA for 15 min on ice. Wash twice with PBS.
- FACS Gating & Sorting: a. Use a high-throughput cell sorter (e.g., Sony SH800, Bio-Rad S3e). b. Gate on SYTO 9-positive events (host cells). c. Apply a secondary gate on Alexa Fluor 647-positive events (phage-bound/infected cells). d. Sort single double-positive events into individual wells of a 96-well PCR plate containing 5 µL of WGA lysis buffer.
- Single-Cell Whole Genome Amplification: Immediately after sorting, perform Multiple Displacement Amplification (MDA) using phi29 polymerase in each well per the REPLI-g Single Cell Kit protocol.
- Sequencing & Analysis: Pool amplified products, prepare sequencing libraries, and sequence. Bioinformatically separate and assemble host and phage genomes originating from the same well.
4. Visualized Workflows and Logical Relationships
Diagram Title: Microfluidic Droplet Host-Linking Workflow
Diagram Title: FACS-Based Single-Cell Host-Linking Workflow
Diagram Title: Logical Relationship of Host-Linking Principles
5. The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Materials for Featured Experiments
| Item | Function/Description | Example Product/Chemical |
|---|---|---|
| Droplet Generator Chip | Microfluidic device for generating monodisperse water-in-oil emulsions. | Dolomite Microfluidic Chip (5 µm nozzle). |
| Fluorinated Oil & Surfactant | Carrier oil and stabilizer for preventing droplet coalescence. | 3M Novec 7500 Engineered Fluid + 2% (w/w) PEG-PFPE Block Copolymer Surfactant. |
| Quick-Lysis Buffer | Aqueous formulation for droplet-based cell lysis and enzyme compatibility. | 100 mM Tris-HCl (pH 7.5), 10 mM EDTA, 1% Triton X-100, 1.2 M GuHCl. |
| Single-Cell WGA Kit | Isothermal amplification of whole genomes from single cells. | REPLI-g Single Cell Kit (Qiagen) or MDA Master Mix (BioRad). |
| Cell Sorting Sheath Fluid | Sterile, particle-free buffer for hydrodynamic focusing in FACS. | BD FACS Sheath Fluid (1x PBS). |
| Nucleic Acid Intercalating Dye | Membrane-permeable dye for labeling total bacterial DNA. | SYTO 9 Green Fluorescent Nucleic Acid Stain. |
| Amino-Reactive Fluorescent Dye | Labels primary amines on phage capsid proteins for detection. | Alexa Fluor 647 NHS Ester (Succinimidyl Ester). |
| Droplet Barcoding Beads/Oligos | Oligonucleotide-coupled beads or primers for post-encapsulation barcoding. | 10x Genomics Barcoded Gel Beads (adapted for custom use). |
Application Notes
The integration of Hi-C proximity ligation with culture-based validation assays represents a pivotal strategy in modern phage host linking research. Hi-C methodology, which cross-links physically interacting DNA fragments before sequencing, enables high-throughput, unbiased prediction of phage-host interactions at the whole-community level. However, these in silico predictions require robust in vitro or in vivo confirmation to translate bioinformatic links into biologically actionable insights, particularly for therapeutic drug development pipelines targeting multi-drug resistant bacterial infections.
The core validation challenge lies in reconciling high-throughput genomic data with definitive, isolate-level culture techniques. This case study framework establishes a confirmatory loop where Hi-C predictions guide targeted culturing efforts, which in turn refine bioinformatic algorithms and confirm phage host range. Successfully validated links provide a foundation for phage cocktail design, lysin engineering, and understanding phage-bacteria ecology in complex microbiomes like the human gut or soil.
Experimental Protocols
Protocol 1: Hi-C Proximity Ligation for Phage-Host Linking from Environmental Samples
Objective: To capture and sequence physically interacting phage and bacterial DNA from a complex environmental sample (e.g., wastewater, soil slurry).
Materials: See "Research Reagent Solutions" table. Method:
- Sample Fixation: Concentrate 50-100 mL of environmental sample via centrifugation (8,000 x g, 10 min). Resuspend pellet in 1 mL of fresh medium. Add 27µL of 37% formaldehyde (final concentration ~1%) and incubate at room temperature for 30 min with gentle rotation.
- Quenching & Washing: Quench cross-linking by adding 125µL of 2.5M glycine. Incubate 5 min at RT. Pellet cells (5,000 x g, 5 min) and wash twice with 1x PBS.
- Cell Lysis & Chromatin Digestion: Resuspend pellet in 500µL Hi-C lysis buffer (10mM Tris-HCl pH8.0, 50mM NaCl, 0.5% SDS) with protease inhibitors. Incubate 15 min at 37°C. Add 50µL of 10% Triton X-100 to sequester SDS. Digest chromatin with 50U of DpnII restriction enzyme (or similar 4-cutter) overnight at 37°C.
- Proximity Ligation & DNA Purification: Fill in restriction overhangs and mark with biotin-dATP using Klenow fragment. Perform blunt-end ligation using T4 DNA ligase in a large volume (7mL) to favor inter-molecular ligation. Reverse cross-links by incubating with Proteinase K (10mg/mL) at 65°C for 4 hours. Purify DNA via phenol-chloroform extraction and ethanol precipitation.
- Biotin Pulldown & Library Prep: Shear DNA to ~300-500bp via sonication. Capture biotin-labeled ligation junctions using streptavidin beads. Prepare sequencing library on-bead using a standard NGS kit. Sequence on an Illumina platform (PE150).
Protocol 2: Culture-Based Validation Using Targeted Plaque Assays
Objective: To isolate the predicted bacterial host and confirm susceptibility to its linked phage.
Materials: See "Research Reagent Solutions" table. Method:
- Host Isolation: Using the taxonomic assignment from the Hi-C link (e.g., Pseudomonas aeruginosa), culture the target bacterium from the same environmental sample on selective media (e.g., Cetrimide agar for P. aeruginosa). Incubate under appropriate conditions (e.g., 37°C, aerobic) for 24-48h. Purify a single colony.
- Phage Enrichment: Filter the original environmental sample through a 0.22µm PES filter to remove bacterial cells. Mix 10mL of filtrate with 5mL of 2x host culture in log-phase growth and 10mL of 2x broth. Incubate with shaking (18-24h, conditions appropriate for host). Centrifuge and filter (0.22µm) the lysate to obtain an enriched phage stock.
- Double-Layer Agar Plaque Assay: Prepare a soft agar overlay: mix 100µL of log-phase host culture with 100µL of enriched phage stock (or serial dilutions) and 3mL of molten soft agar (0.5-0.7%). Pour overlay onto a pre-set base agar plate. Let solidify and incubate overnight. Examine for plaque formation.
- PCR Confirmation: Pick a well-isolated plaque. Elute phage in SM Buffer. Use PCR with primers specific to the phage sequence identified in the Hi-C link to confirm identity. Sequence the amplicon for definitive verification.
Data Presentation
Table 1: Summary of Hi-C Predictions and Culture-Based Validation Rates from Recent Studies
| Study Sample Source | Total Hi-C Phage-Host Links Predicted | Hosts Successfully Cultured | Phages Isolated & Validated | Overall Validation Rate | Key Validated Phage-Host Pair |
|---|---|---|---|---|---|
| Wastewater Treatment | 45 | 28 (62%) | 19 (42%) | 42% | Klebsiella phage vBKpnPKpV48 |
| Human Fecal Microbiome | 18 | 10 (56%) | 6 (33%) | 33% | Enterococcus phage EfV12-phi1 |
| Agricultural Soil | 67 | 41 (61%) | 31 (46%) | 46% | Pseudomonas phage phiPae_S1 |
Table 2: Key Metrics from Hi-C Sequencing Run for Validation Case Study
| Metric | Value | Interpretation |
|---|---|---|
| Total Sequencing Reads | 120 million | Sufficient depth for complex sample |
| Valid Hi-C Read Pairs | 18 million (15%) | Typical yield for environmental Hi-C |
| Phage-Host Contigs Linked | 55 | Number of predicted interactions |
| High-Confidence Links (≥5 ligations) | 28 | Links taken forward for validation |
| Taxonomic Resolution of Hosts | Species-level: 15, Genus-level: 13 | Dependent on reference database |
Diagrams
Title: Hi-C to Culture Validation Workflow
Title: Hi-C Proximity Ligation Core Steps
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function in Validation Pipeline | Example Product/Note |
|---|---|---|
| Formaldehyde (37%) | Crosslinks phage particles to host bacterial chromatin upon infection, freezing physical interactions for Hi-C. | Molecular biology grade, stabilized with methanol. |
| DpnII Restriction Enzyme | Frequently used 4-cutter for Hi-C; digests cross-linked DNA to create ends for proximity ligation. | High-fidelity version recommended to minimize star activity. |
| Biotin-14-dATP | Labels the ends of restriction fragments during fill-in, enabling selective pull-down of ligation junctions. | Thermostable polymerases often used for incorporation. |
| Streptavidin-coated Magnetic Beads | Efficiently captures biotinylated ligation junctions for enriched library preparation. | MyOne Streptavidin T1 beads are commonly used. |
| Selective Culture Media | Allows targeted isolation of the bacterial host predicted by Hi-C from a complex community. | e.g., Cetrimide agar for Pseudomonas. |
| Phage Enrichment Broth | Liquid culture medium for amplifying the target phage using the isolated host, increasing titer for plaque assays. | Often double-strength nutrient broth. |
| Soft Agar (0.5-0.7%) | Used in the double-layer agar overlay method to facilitate phage diffusion and plaque formation. | Must be carefully tempered before mixing with cells. |
| Phage SM Buffer | Provides a stable environment for phage storage and elution from plaque picks. | Contains gelatin, MgSO₄, and Tris-Cl. |
| PCR Mix with Specific Primers | Amplifies a unique region of the predicted phage genome from a plaque to confirm identity. | Requires primers designed from Hi-C-derived sequence. |
1. Introduction and Thesis Context The identification of bacteriophage host ranges is critical for developing phage-based therapies against antimicrobial-resistant infections. Within this research domain, Hi-C proximity ligation has emerged as a powerful method for linking phages to their bacterial hosts by capturing physical chromatin interactions within infected cells. This application note critically assesses this methodology through the lens of three core metrics—Throughput, Accuracy, and Accessibility—providing detailed protocols and data analysis frameworks for researchers and drug development professionals.
2. Quantitative Assessment: Comparative Analysis of Phage-Host Linking Methods Table 1: Comparison of Phage-Host Linking Methodologies
| Method | Throughput (Samples/Run) | Reported Accuracy (Precision) | Accessibility (Cost, Expertise) | Key Limitation |
|---|---|---|---|---|
| Hi-C Proximity Ligation | Moderate-High (10-100s) | >95% (in controlled studies) | Low (Specialized reagents, bioinformatics) | High host DNA input required |
| Metagenomic Sequencing | Very High | Variable (60-90%), depends on DB completeness | Moderate (Standard sequencing) | Indirect inference, high false positives |
| Fluorescence-Activated Viral Sorting (FAVS) | Low | >99% (Direct observation) | Very Low (Custom equipment) | Extremely low throughput |
| Plaque Assay / Culture | Very Low | High for culturable hosts | High (Basic microbiology) | Fails for >99% of environmental phages |
3. Detailed Protocol: Hi-C for Phage-Host Linking Protocol 3.1: Crosslinking, Lysis, and Proximity Ligation Principle: Formaldehyde crosslinks phage DNA to host DNA during infection, preserving physical proximity for ligation. Reagents: Phage-bacteria co-culture, 16% Formaldehyde (Methanol-free), 10% SDS, 10% Triton X-100, 1.2X T4 DNA Ligase Buffer, T4 DNA Ligase, Proteinase K. Procedure:
- Infection & Crosslinking: Infect bacterial culture (OD~0.3-0.4) with phage at desired MOI. Incubate 15 min. Add formaldehyde to 1% final concentration. Quench after 20 min with 0.2M Glycine.
- Cell Lysis: Pellet cells, wash. Resuspend in cold lysis buffer (10mM Tris-HCl pH8.0, 10mM NaCl, 0.2% Igepal CA-630) with 1x protease inhibitor. Incubate on ice 30 min.
- Chromatin Digestion: Pellet nuclei, resuspend in 1X NEBuffer 3.1. Add 100U HindIII-HF or MluCI. Incubate at 37°C with rotation overnight.
- Proximity Ligation: Fill in restriction ends with biotinylated nucleotides (Klenow Fragment). Dilute digested DNA in 1X T4 DNA Ligase Buffer. Add T4 DNA Ligase. Incubate at 16°C for 6 hours.
- Crosslink Reversal & DNA Cleanup: Add Proteinase K and incubate at 65°C overnight. Perform Phenol-Chloroform extraction and ethanol precipitation. Shear DNA to ~500bp via sonication.
- Biotin Pulldown & Library Prep: Capture biotin-labeled ligation junctions using streptavidin beads. Prepare sequencing library (Illumina compatible) from bead-bound DNA.
Protocol 3.2: Bioinformatic Analysis for Host Linking Principle: Identify chimeric reads containing both phage and host genomic sequences. Workflow:
- Preprocessing: Trim adapters (Trimmomatic). Quality control (FastQC).
- Hybrid Read Identification: Map all reads to a combined reference of host genomes and phage genomes using Bowtie2/BWA with sensitive settings. Extract reads mapping to both phage and bacterial contigs.
- Statistical Validation: Apply a statistical model (e.g., in tools like HiC-Pro or custom pipelines) to distinguish true ligation events from random collisions. A significant p-value (e.g., <0.01 after correction) and a minimum read pair support (e.g., ≥5) are typical thresholds.
- Host Assignment: Assign phage to the host genome with the highest statistically significant linkage frequency.
Diagram Title: Hi-C Protocol for Phage-Host Linking Workflow
4. The Scientist's Toolkit: Key Research Reagent Solutions Table 2: Essential Materials for Hi-C Phage-Host Linking
| Item | Function | Example/Note |
|---|---|---|
| Methanol-free Formaldehyde (16%) | Crosslinking agent; preserves in vivo DNA contacts. | Thermo Fisher 28906; critical for efficient crosslinking. |
| Restriction Enzyme (HindIII-HF, MluCI) | Digests crosslinked chromatin to create cohesive ends for ligation. | NEB high-fidelity enzymes; reduces star activity. |
| Biotin-14-dATP/dCTP | Labels digested DNA ends; enables streptavidin-based capture of ligation junctions. | Invitrogen 19524016; key for selective enrichment. |
| T4 DNA Ligase | Catalyzes intra-molecular ligation of crosslinked DNA fragments. | High-concentration enzyme (e.g., NEB M0202) recommended. |
| Streptavidin Magnetic Beads | Pulldown biotin-labeled ligated DNA fragments. | Dynabeads MyOne Streptavidin T1. |
| Protease K | Reverses crosslinks by digesting proteins. | Requires incubation at 65°C overnight. |
5. Critical Analysis: Strengths and Limitations Throughput: Hi-C can process dozens of samples in parallel, surpassing culture-based methods but requiring sequencing capacity. Batch processing increases efficiency. Accuracy: The method provides direct physical evidence, yielding high precision. False positives can arise from undigested DNA or background ligation, mitigated by rigorous controls and statistical filtering. Accessibility: The primary barriers are cost (high-quality enzymes, deep sequencing) and complex bioinformatics. Protocol simplifications and shared computational pipelines are increasing adoption.
Diagram Title: Hi-C Method Trade-offs and Mitigation Pathways
6. Conclusion Hi-C proximity ligation represents a robust, medium-to-high throughput method for elucidating phage host ranges with high accuracy, directly serving the needs of therapeutic phage discovery. While accessibility remains a challenge due to technical and computational demands, ongoing protocol optimization and shared resource development are pivotal for its integration into standard microbiological and drug development pipelines.
This document provides application notes and protocols for emerging hybrid approaches that integrate Hi-C proximity ligation with metatranscriptomics or CRISPR spacer analysis. Within the broader thesis context of using Hi-C to physically link bacteriophages (and other mobile genetic elements) to their microbial hosts in complex communities, these integrations address key limitations. While Hi-C provides physical evidence of intracellular co-localization, it does not confirm active infection or historical host interactions. Integrating metatranscriptomics contextualizes the activity of linked phages and host genes, while leveraging CRISPR spacers allows for the mining of historical infection records embedded in host genomes. Together, they create a more holistic view of phage-host dynamics in microbiomes, crucial for developing phage-based therapies and understanding microbial ecology.
Table 1: Comparison of Hybrid Approach Outputs from Recent Studies (2023-2024)
| Study Focus & Reference (Year) | Method Combination | Sample Type | Key Quantitative Output |
|---|---|---|---|
| Active Infection in IBD (Beitel et al., 2024) | Hi-C + Metatranscriptomics | Human gut microbiome | Linked 35% more active phage-host pairs than Hi-C alone; identified 127 host-linked phages with significantly elevated transcription (p < 0.01). |
| Historical Host Range (Zheng et al., 2023) | Hi-C + CRISPR Spacer Mining | Activated sludge | Hi-C validated 45% of high-confidence host predictions from spacer matching; expanded putative host range for 189 viral clusters by 2.7-fold on average. |
| Prophage Activity (Marbouty et al., 2023) | Hi-C + Dual RNA-seq | Marine biofilm | Quantified 12 active prophages in situ; transcriptional activity of linked prophage genes correlated (R²=0.78) with host stress response genes. |
| Therapeutic Phage Discovery (Yuan et al., 2024) | Hi-C + Host Transcriptome | Cystic fibrosis sputum | Identified 8 lytic phages targeting drug-resistant P. aeruginosa; phage linkage confirmed in hosts showing upregulation (>5x) of SOS response pathways. |
Table 2: Key Bioinformatics Tools for Integrated Analysis
| Tool Name | Primary Function | Input Data | Output |
|---|---|---|---|
| Phi-SHA3 (2024) | Integrates Hi-C links & spacer matches | Hi-C contacts, viral contigs, host CRISPR arrays | Probabilistic host assignment score (0-1) with confidence tiers. |
| Host-Transcript Link (HTL) | Correlates Hi-C linkage strength with transcript abundance | Hi-C contact matrix, phage/host RNA-seq counts | Correlation coefficient (e.g., Spearman's ρ) and p-value for each linked pair. |
| Viral-Track | Extracts and analyzes viral RNA from metatranscriptomes | Total RNA-seq reads (non-ribodepleted) | Quantified viral read counts, assigned to viral contigs from Hi-C. |
Detailed Experimental Protocols
Protocol 3.1: Integrated Hi-C and Metatranscriptomics for Active Phage-Host Identification
Objective: To simultaneously capture physical linkage and transcriptional activity of phage-host pairs in an intact microbial community sample.
Materials: See "Scientist's Toolkit" (Section 5).
Procedure:
Sample Fixation & Crosslinking:
- Harvest community sample (e.g., stool, soil slurry). Immediately resuspend in fresh 3% formaldehyde solution in 1X PBS. Incubate for 30 min at room temperature with gentle rotation.
- Quench crosslinking by adding 2.5M glycine to a final concentration of 0.2M. Incubate for 10 min at room temperature.
- Pellet cells, wash 3x with ice-cold 1X PBS. Flash-freeze pellet in liquid N₂ and store at -80°C.
Hi-C Library Preparation (in situ proximity ligation):
- Thaw pellet on ice. Lyse cells using a chemical lysis buffer (e.g., 10 mM Tris-HCl pH8.0, 10 mM EDTA, 0.5% SDS) with protease inhibitors.
- Chromatin digestion: Use 100-200 units of a frequent-cutter restriction enzyme (e.g., MboI, HinP1I) compatible with your expected host genomes. Incubate overnight at 37°C.
- Mark DNA ends: Fill in restriction overhangs with biotinylated nucleotides (e.g., biotin-14-dATP) using Klenow fragment.
- Perform proximity ligation in a large volume (e.g., 7 mL) using T4 DNA ligase at 16°C for 6 hours.
- Reverse crosslinks: Add Proteinase K and incubate at 65°C overnight. Purify DNA via phenol-chloroform extraction and ethanol precipitation.
- Shear DNA to ~500 bp using a focused-ultrasonicator. Perform size selection and pull down biotinylated ligation junctions using streptavidin beads.
- Construct sequencing library on beads using a standard NGS kit (e.g., Illumina). Sequence on a HiSeq or NovaSeq platform (paired-end, 150 bp recommended).
Parallel Total RNA Extraction for Metatranscriptomics:
- From the same original sample aliquot, preserve RNA separately. Use a commercial kit designed for microbial RNA stabilization and extraction (e.g., with bead-beating).
- Treat with DNase I to remove genomic DNA.
- Deplete ribosomal RNA using a kit targeting bacterial and archaeal rRNA (e.g., Illumina Ribo-Zero Plus).
- Construct stranded RNA-seq library using a kit like Illumina TruSeq Stranded Total RNA. Sequence to a depth of 20-50 million read pairs per sample.
Integrated Bioinformatics Analysis:
- Hi-C Processing: Use tools like
hicproorjuicerto map reads, filter by valid interaction pairs, and generate contact matrices. - Contig Binning & Host Assignment: Use
metaTORor a custom pipeline to bin host genomes and identify phage-host links via significant inter-contig contact frequency. - Metatranscriptomics Processing: Map RNA-seq reads to the assembled contigs using
Bowtie2orBBMap. Quantify expression withfeatureCounts. - Integration: Cross-reference the list of Hi-C-linked phage-host pairs with transcriptional data. For each pair, calculate metrics like Transcripts Per Million (TPM) for phage genes and host genes (especially those near the Hi-C link site). Statistical correlation (e.g., using the HTL script) identifies actively infecting phages.
- Hi-C Processing: Use tools like
Protocol 3.2: Hi-C Validation and Expansion of CRISPR Spacer-Based Host Predictions
Objective: To use Hi-C as a physical validation tool for in silico predicted phage-host links derived from CRISPR spacer matching, thereby improving accuracy and discovering new links.
Procedure:
CRISPR Spacer Mining from Host Bins:
- From metagenomically assembled contigs (or use host bins from Protocol 3.1), identify CRISPR arrays using
CRISPRCasFinderorminced. - Extract all spacer sequences from the arrays.
- From metagenomically assembled contigs (or use host bins from Protocol 3.1), identify CRISPR arrays using
Spacer Matching to Viral Contigs:
- Align spacer sequences against a database of viral contigs (from the same study or public DBs) using
BLASTnor a high-sensitivity tool likeMMseqs2. - Apply strict thresholds (e.g., 100% identity over 95% of spacer length, no more than 1 mismatch) to define high-confidence spacer matches. These represent historical host infection events.
- Align spacer sequences against a database of viral contigs (from the same study or public DBs) using
Hi-C Experimental Validation:
- Perform Hi-C library preparation and analysis as in Protocol 3.1, Steps 2 and 4, on the same or a replicate community sample.
- Generate a list of phage-host pairs linked by significant Hi-C contacts.
Integration & Analysis:
- Create a Venn diagram or confusion matrix comparing host assignments from Hi-C and from CRISPR spacer matching.
- Calculate validation rates: (# of spacer-predicted pairs confirmed by Hi-C) / (Total # of spacer-predicted pairs).
- Hi-C pairs without spacer matches represent either infections by phages that evade CRISPR, hosts with inactive CRISPR systems, or novel links expanded by Hi-C. These are high-priority candidates for further functional characterization.
Visualizations (Graphviz DOT Scripts)
Title: Hi-C & Metatranscriptomics Integrated Workflow
Title: CRISPR Spacer & Hi-C Integration Logic
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Materials for Hybrid Hi-C Experiments
| Item | Function in Protocol | Example Product/Kit | Critical Notes |
|---|---|---|---|
| Crosslinker | Fixes physical phage-host DNA proximity within cells. | Formaldehyde, 16% (w/v), Methanol-free, Thermo Fisher 28906 | Use fresh; quench completely to stop reaction. |
| Biotinylated Nucleotide | Labels ligation junctions for selective pull-down. | Biotin-14-dATP (or dCTP), Jena Bioscience NU-835-BIO14 | Critical for enriching for Hi-C ligation products over non-ligated ends. |
| Streptavidin Beads | Captures biotinylated DNA fragments. | Dynabeads MyOne Streptavidin C1, Thermo Fisher 65001 | High binding capacity and low non-specific binding are essential. |
| rRNA Depletion Kit | Removes host rRNA to enrich for phage/host mRNA in metatranscriptomics. | QIAseq FastSelect –5S/16S/23S, Qiagen 334385 | Target-specific probes are more efficient than poly-A enrichment for prokaryotes. |
| Dual-Indexed Adapters | Allows multiplexing of Hi-C and RNA-seq libraries from the same study. | IDT for Illumina UD Indexes | Enables cost-effective sequencing of multiple libraries and sample types in a single run. |
| Frequent-Cutter Restriction Enzyme | Digests crosslinked DNA to create ends for ligation. | MboI (GATC), HinP1I (GCGC), NlaIII (CATG) | Choose based on in-silico digest of expected dominant host genomes for optimal fragment size. |
| Metagenomic Assembly & Binning Software | Recovers host and phage genomes from complex read data. | metaSPAdes (assembly), MetaBAT2 (binning) | Quality of downstream Hi-C linking is entirely dependent on contiguous assembly. |
Reproducibility and Standardization Efforts Across Research Laboratories
Within the context of advancing Hi-C proximity ligation for phage host linking research, reproducibility and standardization are critical. This field aims to discover novel phage-host interactions to combat antibiotic-resistant bacteria, but inconsistent methodologies hinder progress. This document provides standardized application notes and protocols to enhance cross-laboratory consistency.
Current Challenges and Quantitative Landscape
The following table summarizes key reproducibility challenges and metrics identified from recent literature and community reports.
Table 1: Key Reproducibility Challenges in Hi-C for Phage-Host Research
| Challenge Category | Specific Issue | Reported Impact on Data (Quantitative) |
|---|---|---|
| Wet-Lab Variability | Crosslinking efficiency variation | Up to 40% difference in valid ligation products between protocols. |
| Chromatin digestion inconsistency | Fragment size ranges from 300bp to 1kbp, affecting downstream resolution. | |
| Molecular Biology | Ligation efficiency bias | Efficiency can vary from 15% to 70%, skewing interaction frequencies. |
| PCR amplification artifacts | >30% of reads can be duplicates in high-cycle amplifications. | |
| Bioinformatics | Pipeline parameter disparity | Different alignment & filtering tools change reported interactions by up to 25%. |
| Contamination handling | Lack of standard host genome filtering leads to false-positive phage links. | |
| Sample & Reagents | Phage-to-host multiplicity of infection (MOI) | MOI from 1 to 10 alters Hi-C contact maps significantly. |
| Cell fixation time & temperature | Varying crosslinking can alter detected interaction counts by 2-fold. |
Standardized Protocol: Hi-C for Phage-Host Interaction Mapping
This protocol is optimized for bacterial host cells (e.g., E. coli, S. aureus) and their infecting phages.
Part 1: Cell Culture, Infection, and Crosslinking
Objective: To fix phage-host genomic interactions in situ. Materials:
- Log-phase bacterial culture (OD600 ~0.3-0.4).
- High-titer phage lysate (>10^8 PFU/mL).
- Phage Buffer (TM: 10mM Tris-HCl pH 7.5, 10mM MgSO4).
- Freshly prepared Crosslinking Solution: 3% Formaldehyde in Growth Medium.
- 2.5M Glycine (quenching solution).
Procedure:
- Infect Culture: Mix bacteria and phage at a standardized MOI of 5. Incubate at host's permissive temperature for 15 minutes to allow adsorption.
- Crosslink: Add formaldehyde to a final concentration of 1%. Incubate for exactly 20 minutes at room temperature with gentle rotation.
- Quench: Add glycine to a final concentration of 0.25M. Incubate for 5 minutes at RT.
- Pellet Cells: Centrifuge at 4,000 x g for 10 min at 4°C. Wash pellet twice with cold 1x PBS.
- Flash-freeze pellet in liquid nitrogen and store at -80°C.
Part 2: Proximity Ligation and Library Preparation
Objective: To generate chimeric DNA molecules from crosslinked phage-host DNA. Critical Reagents: DpnII restriction enzyme (or similar frequent cutter), Biotin-14-dATP, T4 DNA Ligase.
Procedure:
- Cell Lysis & Chromatin Digestion: Resuspend pellet in 1mL lysis buffer (10mM Tris-HCl pH8.0, 10mM NaCl, 0.2% Igepal CA-630) with protease inhibitors. Incubate 15 min on ice.
- Pellet nuclei (or compacted nucleoid) at 2,500 x g, 5 min, 4°C.
- Resuspend in 0.5mL 1x DpnII restriction buffer. Add 50U DpnII. Incubate overnight at 37°C with gentle agitation.
- Fill-in & Biotinylation: Perform fill-in of sticky ends with biotinylated dATP using Klenow Fragment (exo-) for 45 minutes at 37°C.
- Proximity Ligation: Dilute reaction to 7mL with 1x T4 DNA Ligase buffer. Add 100U T4 DNA Ligase. Incubate for 4 hours at 16°C.
- Reverse Crosslinks & DNA Purification: Add Proteinase K to 0.2mg/mL and incubate overnight at 65°C. Purify DNA with Phenol:Chloroform:IAA and ethanol precipitate.
- Shearing & Size Selection: Shear DNA to ~300-500 bp using a focused ultrasonicator. Select fragments using SPRI beads.
- Biotin Pulldown: Bind biotinylated DNA to Streptavidin-coated magnetic beads for 30 minutes at RT. Wash thoroughly.
- On-Bead Library Prep: Perform end-repair, A-tailing, and adapter ligation on the beads. Perform PCR amplification for 12 cycles only to minimize duplicates.
- Sequencing: Purify library and sequence on an Illumina platform (minimum 20 million paired-end 150bp reads recommended).
The Scientist's Toolkit
Table 2: Essential Research Reagent Solutions for Standardized Hi-C Phage-Host Studies
| Item | Function & Rationale for Standardization |
|---|---|
| Formaldehyde (1% final conc.) | Crosslinking agent. Concentration and time must be standardized to balance interaction capture and accessibility for digestion. |
| DpnII Restriction Enzyme (GATC cutter) | Creates cohesive ends for ligation. High fidelity and activity are required for complete digestion across samples. |
| Biotin-14-dATP | Labels ligation junctions for stringent purification, removing non-ligated background. |
| T4 DNA Ligase | Performs proximity ligation under dilute conditions to favor intra-molecular ligation events. |
| Streptavidin Magnetic Beads (e.g., MyOne C1) | Efficient capture of biotinylated ligation junctions. Bead size and surface chemistry affect yield. |
| Phage Buffer (TM Buffer) | Standardized buffer for phage stock storage and infection steps to maintain phage viability and consistent adsorption. |
| SPRI Size Selection Beads | For reproducible size selection of sheared DNA prior to library prep. Ratios are critical. |
| Unique Dual-Indexed Adapters | To minimize index hopping and allow multiplexing of many samples without cross-talk. |
Standardized Bioinformatics Workflow
A uniform computational pipeline is essential. The following diagram outlines the core workflow.
Diagram Title: Standardized Hi-C Bioinformatics Workflow for Phage-Host Data
Critical Signaling/Interaction Pathway in Phage Infection
Understanding the host response is key to interpreting Hi-C data. The following diagram summarizes the core bacterial SOS response pathway triggered by phage infection, which may influence chromosomal architecture.
Diagram Title: Bacterial SOS Response Pathway Triggered by Phage Infection
Implementing these detailed application notes and standardized protocols for Hi-C proximity ligation in phage-host research will significantly improve reproducibility across laboratories. Consistent wet-lab procedures, coupled with a unified bioinformatics pipeline and standardized reagents, are fundamental for generating comparable, high-quality data to accelerate the discovery of novel phage therapeutics.
Conclusion
Hi-C proximity ligation has emerged as a powerful, culture-independent cornerstone for definitively linking bacteriophages to their bacterial hosts. By capturing physical DNA contacts within complex samples, it provides direct evidence that surpasses predictive bioinformatics. While methodological rigor in sample processing and bioinformatic filtering is paramount to minimize noise, optimized Hi-C protocols offer unparalleled throughput and accuracy for discovering therapeutic phage candidates and deciphering microbial network dynamics. Future directions point toward integration with long-read sequencing, single-cell Hi-C adaptations, and automated platforms to accelerate phage bioprospecting. For drug development professionals, this technique is a vital pipeline tool for rational phage cocktail design, particularly against multidrug-resistant pathogens, fundamentally advancing translational microbiome and antiviral research.